work in progress...
About
Walkthrough: How to use FsFast and fcseed-sess for functional connectivity analysis including example commands.
For general tips on using FsFast, download this FS-FAST powerpoint
This walkthrough is based on
*STEP 1: Unpack Data into the FSFAST Hierarchy using unpacksdcmdir
Ex: unpacksdcmdir -src dicomdir/subject/ALLDICOMS -targ fcMRI_dir/subject -cfg subject_config.txt -fsfast -unpackerr
In this example command...
- Have all fMRI dicoms linked into "ALLDICOMS" directory
- Arguement for "-targ" specifies output directory
- subject_config.txt is a configuration text file you create (format below)
- Use "-fsfast" to generate fsfast hierarchy
subject_config.txt format:
28 bold nii f.nii 29 bold nii f.nii
Col.1: scan acquisition number Col.2: output dir name will be created within "fcMRI_dir/subject" Col.3: output file format - this example is nifti format Col.4: output filename. In this example, 2 files will be created:
- fcMRI_dir/subject/028/f.nii fcMRI_dir/subject/029/f.nii
1.QA Check after unpacking:
- A - Check unpacked data (time points, # of slices ..etc)
- B - Check FSFAST hierarchy in session folder
*STEP 2: Reconstruction Anatomical data using recon-all
Ex:
setenv SUBJECTS_DIR /path/to/recon_dir/ recon-all -s subject_dirname -all -i pathtoT1dicom_scan1.dcm -i pathtoT1dicom_scan2.dcm
In this example command...
set your SUBJECTS_DIR variable to your FreeSurfer subject recon directory
- set the subject's directory name with "-s" ... the arguement you provide will become the directory name within $SUBJECTS_DIR
- use "-i" to supply the dicoms to reconstruct. Use one "-i" per T1 acquisition.
2.QA Check:
- A - Check talairach transformation
B - Check skull strip, white matter & pial surface
- C - Re-run "recon-all" if edits are made
- D - Check hierarchy of reconstructed anatomical data
1.Make FSFAST basic hierarchy (only if data are not unpacked in FSFAST hierarchy)
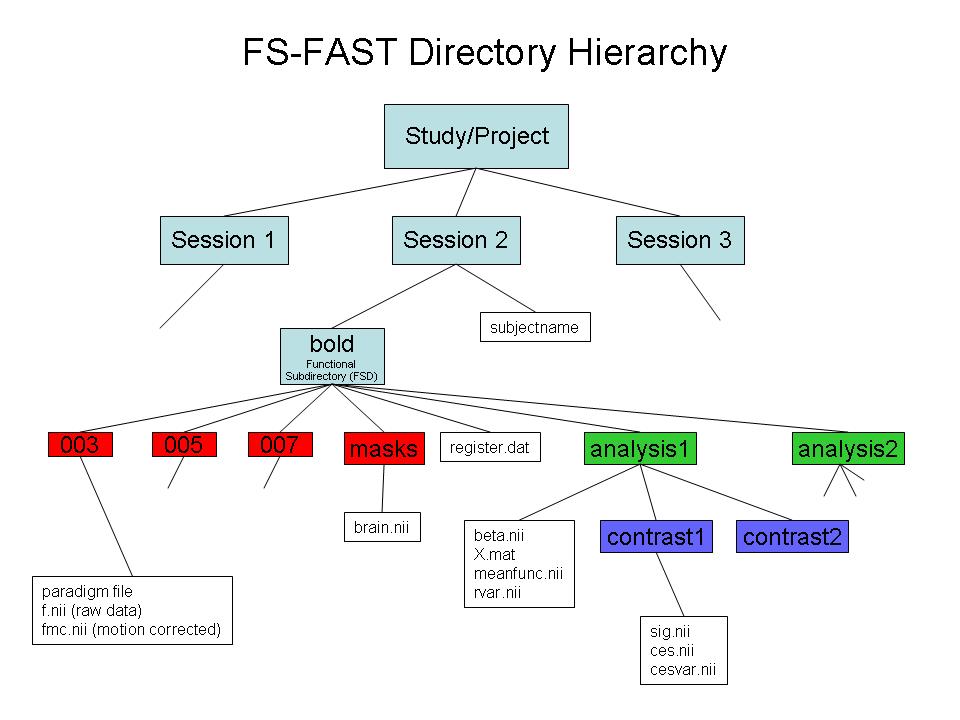
2.Link to FreeSurfer anatomical analysis
A - Make subjectname file in the session directory to link a subject's functional & structural data
3.Create a sessid file (text file with list of your sessions)in your Study DIR.
4.Create a Stimulus Schedule (Paradigm file) in bold folder (A "paradigm" file is a record of which stimulus was presented when & for how long.
Each paradigm file has four columns:
- A - Stimulus onset time (sec)
- B - Condition ID code (0, 1, 2, ...)
- C - Stimulus Duration (sec)
- D - Stimulus Weight (usually 1)
*STEP 3: Pre-process your bold data using preproc-sess preproc-sess
# Preprocessing of fMRI Data
preproc-sess -s <subjid> -fwhm <#>
1.By default this will do motion correction, smoothing & brain masking
2.Quality Check (plot-twf-sess) 3.Examine additions to FSFAST hierarchy (in each run of bold dir):
- f.nii (Raw fMRI data)
- fmc.nii (Motion corrected-MC)
fmcsm5.nii (MC & smoothed)
- fmc.mcdat (Text file with the MC parameters (AFNI))
- brain.mgz (Binary mask of the brain)
# Function-Structure Registration View unregistered:
tkregister-sess -s <subjid> -regheader)
Run automatic registration:
spmregister-sess -s <subjid>
Check automatic registration:
tkregister-sess -s <subjid>
A - Make edits if needed using scale as the last resort Check talairach registration:
tkregister2 --s <subjid> --fstal --surf
*STEP 4: Use fcseed-sess to generate time-course information for your chosen seed region (as well as nuisance variable signal).
*STEP 5: Use mkanalysis-sess to setup an analysis for your FC data
*STEP 6: Use selxavg3-sess to run the subject-level analysis
*STEP 7: Use mri_glmfit or selxavg3-sess to run a group-level analysis
