|
Size: 19301
Comment:
|
Size: 23078
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 32: | Line 32: |
| # generating_program /usr/local/freesurfer/stable6/bin/dmri_pathstats # cvs_version # cmdline /usr/local/freesurfer/stable6/bin/dmri_pathstats --intrc /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr --dtbase /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dmri/dtifit --path lh.ilf --subj elmo.2012 --out /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr/pathstats.overall.txt --outvox /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr/pathstats.byvoxel.txt |
# generating_program /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats # cvs_version ayendiki-local # cmdline /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats --intrc /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr --dtbase /space/erebus/1/users/data/trc/subject1/dmri/dtifit --path lh.ilf --subj subject1 --invox path.ref.txt --out /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt --outvox /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.byvoxel.txt |
| Line 36: | Line 36: |
| # hostname oribi | # hostname compute-0-124.nmr.mgh.harvard.edu |
| Line 38: | Line 38: |
| # user eboyd2 | # user ayendiki |
| Line 41: | Line 41: |
| # subjectname elmo.2012 | # subjectname subject |
| Line 45: | Line 45: |
| Volume 629 Len_Min 36 Len_Max 71 Len_Avg 50.01 Len_Center 48 AD_Avg 0.00136398 AD_Avg_Weight 0.00137336 AD_Avg_Center 0.00137163 RD_Avg 0.000570444 RD_Avg_Weight 0.000568755 RD_Avg_Center 0.00056212 MD_Avg 0.000834957 MD_Avg_Weight 0.000836957 MD_Avg_Center 0.000831957 FA_Avg 0.505701 FA_Avg_Weight 0.510267 FA_Avg_Center 0.514496 |
Volume 1996 Len_Min 53 Len_Max 129 Len_Avg 79.9787 Len_Center 89 AD_Avg 0.00120044 AD_Avg_Weight 0.00121209 AD_Avg_Center 0.00123318 RD_Avg 0.000467543 RD_Avg_Weight 0.000458132 RD_Avg_Center 0.000549716 MD_Avg 0.000711842 MD_Avg_Weight 0.000709453 MD_Avg_Center 0.000777538 FA_Avg 0.540167 FA_Avg_Weight 0.553683 FA_Avg_Center 0.477498 |
| Line 134: | Line 134: |
| # generating_program /usr/local/freesurfer/stable6/bin/dmri_pathstats # cvs_version # cmdline /usr/local/freesurfer/stable6/bin/dmri_pathstats --intrc /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr --dtbase /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dmri/dtifit --path lh.ilf --subj elmo.2012 --out /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr/pathstats.overall.txt --outvox /autofs/space/rama_001/users/eboyd2/FStutorial_edits/diffusion_tutorial/elmo.2012/dpath/lh.ilf_AS_avg33_mni_bbr/pathstats.byvoxel.txt |
# generating_program /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats # cvs_version ayendiki-local # cmdline /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats --intrc /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr --dtbase /space/erebus/1/users/data/trc/subject1/dmri/dtifit --path lh.ilf --subj subject1 --invox path.ref.txt --out /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt --outvox /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.byvoxel.txt |
| Line 138: | Line 138: |
| # hostname oribi | # hostname compute-0-124.nmr.mgh.harvard.edu |
| Line 140: | Line 140: |
| # user eboyd2 | # user ayendiki |
| Line 143: | Line 143: |
| # subjectname elmo.2012 | # subjectname subject1 |
| Line 148: | Line 148: |
| 83 74 10 0.0010087 0.000830648 0.000889999 0.159232 0.00116397 0.000637616 0.000813066 0.374542 83 73 10 0.00107542 0.00076142 0.000866087 0.248381 0.00116 0.000631537 0.000807689 0.378022 83 73 11 0.00110213 0.000712833 0.000842598 0.275052 0.00116456 0.000632124 0.000809599 0.378441 83 72 11 0.00115836 0.000627003 0.000804122 0.379383 0.00118313 0.000638984 0.000820367 0.378685 83 71 11 0.00130141 0.00050243 0.000768757 0.540997 0.00120207 0.000613241 0.000809521 0.409762 83 71 12 0.00135934 0.000596314 0.000850654 0.479475 0.00122036 0.00060423 0.00080961 0.425146 83 70 12 0.00154584 0.000596379 0.000912867 0.540192 0.0012735 0.000598274 0.000823352 0.449849 83 69 12 0.0015961 0.000637489 0.000957027 0.523057 0.0013166 0.000627619 0.000857281 0.443408 83 68 13 0.00144106 0.000659587 0.000920077 0.456897 0.00131143 0.000646806 0.000868347 0.428068 84 67 13 0.00133104 0.000563732 0.000819502 0.501266 0.00126369 0.00063388 0.000843816 0.423312 84 66 14 0.00129191 0.000639349 0.000856869 0.429424 0.00120567 0.000636186 0.000826019 0.401576 84 65 14 0.00111743 0.000576156 0.000756579 0.401439 0.00120671 0.000675135 0.000852329 0.370139 84 64 15 0.000995878 0.000653256 0.000767463 0.27065 0.00115213 0.000677309 0.000835582 0.347765 84 63 15 0.00103649 0.000619989 0.000758823 0.333934 0.00120864 0.000672818 0.00085142 0.372466 84 62 16 0.00121678 0.000586189 0.000796385 0.432171 0.00128202 0.000629584 0.000847063 0.436846 84 61 16 0.00128539 0.000496937 0.000759754 0.553296 0.00133043 0.000553918 0.000812759 0.516887 84 60 16 0.00128823 0.00053104 0.000783438 0.53335 0.00134452 0.000506174 0.000785624 0.563432 84 60 17 0.00126802 0.000460207 0.000729479 0.58859 0.00133849 0.000496096 0.000776899 0.570689 84 59 17 0.00108238 0.000373869 0.000610039 0.629474 0.00133839 0.000466135 0.00075689 0.597466 84 58 17 0.00130795 0.000494729 0.000765803 0.58186 0.00140401 0.000490494 0.000795002 0.591723 84 57 18 0.00107557 0.000464563 0.000668231 0.525115 0.00141309 0.000503638 0.000806786 0.580816 84 56 18 0.00112712 0.000392396 0.000637304 0.612159 0.00142693 0.000501972 0.000810288 0.593075 84 55 19 0.00110705 0.000536791 0.000726878 0.47699 0.0014481 0.000521233 0.000830186 0.580617 84 54 19 0.00131515 0.000501804 0.000772919 0.560854 0.00149121 0.000514767 0.000840246 0.591875 84 53 19 0.00141376 0.000488109 0.000796661 0.592974 0.00151916 0.00050704 0.000844416 0.603714 84 53 20 0.00119967 0.000528667 0.000752334 0.515978 0.00151852 0.000501309 0.000840379 0.609157 84 52 20 0.00120838 0.000496335 0.000733684 0.545742 0.00151907 0.000506619 0.0008441 0.607991 84 51 21 0.0011066 0.000545991 0.000732861 0.48208 0.00145287 0.000537145 0.000842389 0.566021 83 50 21 0.00148025 0.000517207 0.000838221 0.591659 0.00143009 0.000558962 0.00084934 0.541725 83 49 21 0.00144729 0.000608569 0.000888142 0.504581 0.00143929 0.000580943 0.000867057 0.523038 83 48 21 0.00140066 0.000648933 0.000899507 0.45017 0.00146562 0.000588808 0.000881079 0.52329 82 47 22 0.00179437 0.000560765 0.000971966 0.632207 0.00152193 0.000564541 0.00088367 0.560292 82 46 22 0.00140457 0.000546434 0.00083248 0.540164 0.00148664 0.000549511 0.000861887 0.56362 81 45 22 0.00165706 0.000706209 0.00102316 0.494609 0.00148319 0.000559383 0.000867323 0.558307 80 44 23 0.00174696 0.000418429 0.000861272 0.723202 0.0014574 0.00058681 0.000877008 0.532129 80 43 23 0.00146997 0.000619092 0.000902717 0.497525 0.00140823 0.000612242 0.000877576 0.493454 79 42 23 0.00152992 0.000444628 0.000806391 0.661404 0.00139555 0.000603363 0.000867423 0.492408 78 41 23 0.00157958 0.000555758 0.000897033 0.592812 0.00138554 0.0005864 0.000852775 0.498505 78 40 24 0.00169536 0.000341967 0.000793098 0.769893 0.00136858 0.00057877 0.000842035 0.494142 77 39 24 0.00176535 0.000454217 0.000891263 0.699291 0.00136671 0.000564577 0.000831952 0.504254 76 39 24 0.00181275 0.000391344 0.000865146 0.750689 0.00138579 0.00055527 0.000832107 0.517531 76 38 25 0.00177161 0.000446334 0.000888092 0.705912 0.00138728 0.000545684 0.000826213 0.523804 75 37 25 0.00186546 0.000517308 0.000966693 0.673821 0.0013714 0.000562255 0.000831969 0.506898 74 36 25 0.00161531 0.000582424 0.000926721 0.575432 0.00135436 0.000571873 0.000832701 0.492641 74 35 25 0.00158317 0.000631253 0.000948557 0.529855 0.00134491 0.000573929 0.000830925 0.489379 73 34 25 0.00146608 0.000741013 0.000982702 0.410368 0.00134846 0.000580893 0.000836751 0.486019 72 34 25 0.00118521 0.000661327 0.000835955 0.367785 0.00134222 0.000586273 0.000838257 0.478949 72 33 25 0.00120423 0.000714338 0.000877636 0.354421 0.00133965 0.000594724 0.000843032 0.47204 |
97 78 16 0.000867977 0.000725794 0.000773188 0.108892 0.00107168 0.000599572 0.000756942 0.363722 97 78 16 0.000867977 0.000725794 0.000773188 0.108892 0.00107168 0.000599572 0.000756942 0.363722 97 77 16 0.000832794 0.000712196 0.000752395 0.098711 0.00106892 0.000599915 0.000756251 0.363217 97 77 16 0.000832794 0.000712196 0.000752395 0.098711 0.00106892 0.000599915 0.000756251 0.363217 96 77 17 0.000778157 0.000713259 0.000734892 0.0524794 0.00106822 0.000599456 0.000755712 0.364947 96 76 17 0.000872096 0.000706771 0.000761879 0.136309 0.00106737 0.00059909 0.000755185 0.365999 96 76 17 0.000872096 0.000706771 0.000761879 0.136309 0.00106737 0.00059909 0.000755185 0.365999 96 75 18 0.00106175 0.000671836 0.000801806 0.326543 0.00106621 0.000593117 0.000750814 0.370047 96 74 19 0.00084758 0.000742526 0.000777544 0.146359 0.00107689 0.000564038 0.000734991 0.399445 96 74 19 0.00084758 0.000742526 0.000777544 0.146359 0.00107689 0.000564038 0.000734991 0.399445 96 73 20 0.00105676 0.000582545 0.000740616 0.379989 0.00109462 0.000529155 0.000717645 0.442998 96 73 20 0.00105676 0.000582545 0.000740616 0.379989 0.00109462 0.000529155 0.000717645 0.442998 96 73 21 0.00106829 0.000535296 0.000712959 0.446277 0.00110578 0.000510944 0.000709221 0.466119 96 73 21 0.00106829 0.000535296 0.000712959 0.446277 0.00110578 0.000510944 0.000709221 0.466119 96 72 21 0.00112798 0.000524413 0.000725602 0.490875 0.00109594 0.000500484 0.000698966 0.476156 96 72 22 0.00100371 0.000518413 0.00068018 0.466196 0.00109198 0.000491746 0.000691821 0.484992 96 71 22 0.00110846 0.000516744 0.000713983 0.496827 0.00107691 0.000488262 0.000684476 0.487281 96 70 23 0.000963374 0.000468995 0.000633788 0.515347 0.00103651 0.000500421 0.000679117 0.471307 96 70 24 0.000952905 0.00052878 0.000670155 0.464126 0.00102183 0.000507845 0.000679174 0.463955 96 69 25 0.000941334 0.000510755 0.000654282 0.452879 0.0010153 0.000520488 0.000685426 0.451515 96 69 25 0.000941334 0.000510755 0.000654282 0.452879 0.0010153 0.000520488 0.000685426 0.451515 96 68 25 0.000937902 0.000498823 0.000645183 0.473998 0.0010151 0.000521116 0.000685777 0.45532 96 67 26 0.00114779 0.000422178 0.00066405 0.597208 0.00106285 0.000495767 0.000684795 0.488664 96 67 26 0.00114779 0.000422178 0.00066405 0.597208 0.00106285 0.000495767 0.000684795 0.488664 96 66 27 0.00119928 0.000456432 0.000704049 0.567628 0.00113942 0.000464739 0.000689633 0.535918 96 66 27 0.00119928 0.000456432 0.000704049 0.567628 0.00113942 0.000464739 0.000689633 0.535918 96 65 27 0.00141769 0.000388493 0.000731558 0.691121 0.00118206 0.000446907 0.000691963 0.56636 96 64 28 0.00139466 0.000325184 0.000681676 0.734954 0.00123269 0.000410431 0.000684517 0.612061 96 64 28 0.00139466 0.000325184 0.000681676 0.734954 0.00123269 0.000410431 0.000684517 0.612061 96 63 28 0.00143303 0.000312452 0.000685979 0.752811 0.00126731 0.000393664 0.000684881 0.636549 95 62 29 0.00130884 0.000290893 0.000630209 0.745983 0.00129524 0.000387733 0.000690235 0.648474 94 62 29 0.00164671 0.00076911 0.00106164 0.446289 0.00129674 0.000390088 0.000692304 0.647809 94 61 29 0.00172456 0.000745308 0.00107172 0.485371 0.00127988 0.000398742 0.000692451 0.638261 94 60 30 0.00140745 0.000880179 0.00105594 0.281852 0.00125246 0.000428793 0.00070335 0.606134 94 60 30 0.00140745 0.000880179 0.00105594 0.281852 0.00125246 0.000428793 0.00070335 0.606134 94 59 31 0.00155751 0.000862578 0.00109422 0.356395 0.00121421 0.000460178 0.000711525 0.56957 94 58 32 0.00130825 0.000701944 0.000904047 0.376433 0.00120559 0.000462724 0.00071035 0.562654 94 58 32 0.00130825 0.000701944 0.000904047 0.376433 0.00120559 0.000462724 0.00071035 0.562654 94 57 33 0.00127009 0.000453103 0.000725431 0.580671 0.001259 0.000436023 0.000710351 0.597881 94 56 34 0.00128442 0.000408392 0.0007004 0.624911 0.00131045 0.000409303 0.000709684 0.635395 94 56 34 0.00128442 0.000408392 0.0007004 0.624911 0.00131045 0.000409303 0.000709684 0.635395 94 55 34 0.00129227 0.00046328 0.00073961 0.577816 0.0013183 0.000405609 0.000709842 0.640691 94 54 35 0.0012703 0.000405959 0.000694073 0.624761 0.00132773 0.000405781 0.000713098 0.639696 94 54 36 0.0012825 0.000416241 0.000704992 0.62111 0.00131601 0.00041376 0.000714509 0.62888 94 54 36 0.0012825 0.000416241 0.000704992 0.62111 0.00131601 0.00041376 0.000714509 0.62888 93 53 36 0.00139747 0.000604594 0.000868887 0.488666 0.00130335 0.000422725 0.000716265 0.616271 93 52 36 0.00135543 0.000585794 0.000842337 0.488772 0.0012934 0.000424784 0.000714323 0.612202 93 52 37 0.00123236 0.000470209 0.000724259 0.552723 0.00128567 0.000433751 0.000717722 0.600983 93 51 38 0.00114102 0.000441359 0.00067458 0.550956 0.00128498 0.000448106 0.000727063 0.587959 93 50 38 0.00115213 0.00050062 0.000717791 0.500001 0.00128811 0.00045668 0.00073382 0.580909 93 50 39 0.00106929 0.000415801 0.000633631 0.546817 0.00129122 0.00045262 0.000732149 0.585613 92 49 39 0.00139456 0.000653969 0.000900833 0.448212 0.00129614 0.000456588 0.000736437 0.583877 92 48 39 0.00121658 0.000614545 0.000815223 0.410542 0.00129869 0.00045225 0.000734397 0.588016 92 48 39 0.00121658 0.000614545 0.000815223 0.410542 0.00129869 0.00045225 0.000734397 0.588016 92 47 40 0.00118115 0.000483847 0.000716281 0.521239 0.0013064 0.00044644 0.000733091 0.593569 91 46 41 0.00132381 0.000679848 0.000894501 0.400171 0.00130968 0.000445202 0.000733361 0.593125 91 46 41 0.00132381 0.000679848 0.000894501 0.400171 0.00130968 0.000445202 0.000733361 0.593125 91 45 41 0.00113019 0.000529645 0.000729826 0.454707 0.001325 0.000431276 0.000729183 0.607594 91 44 41 0.00117487 0.000464237 0.000701113 0.530762 0.00134519 0.00041734 0.000726622 0.623841 90 43 41 0.00121382 0.000492301 0.000732807 0.519596 0.00135435 0.00040553 0.000721804 0.635504 90 42 42 0.00141832 0.000419311 0.000752312 0.650028 0.00135835 0.000396281 0.000716967 0.644296 90 41 43 0.00149731 0.000356095 0.0007365 0.72266 0.00135424 0.000393535 0.00071377 0.64191 89 40 43 0.001381 0.000513018 0.000802344 0.564421 0.00132657 0.000407598 0.000713924 0.624407 89 40 43 0.001381 0.000513018 0.000802344 0.564421 0.00132657 0.000407598 0.000713924 0.624407 90 39 43 0.00157585 0.000295255 0.000722119 0.787805 0.00130086 0.000413205 0.00070909 0.613173 89 38 43 0.00138324 0.000383207 0.000716551 0.674917 0.00125725 0.000434189 0.000708544 0.581023 89 38 44 0.00124692 0.000510601 0.000756042 0.514663 0.001253 0.000438098 0.000709733 0.575357 89 37 45 0.00118923 0.00056238 0.000771331 0.44532 0.00122624 0.00044834 0.000707639 0.557656 88 37 45 0.00109356 0.000633894 0.000787115 0.365015 0.00123292 0.000451307 0.000711844 0.55797 88 36 45 0.00103814 0.000676695 0.000797177 0.316449 0.00120901 0.000458568 0.000708718 0.544602 88 35 45 0.00114301 0.000589147 0.000773767 0.391688 0.00118172 0.000467419 0.000705519 0.529077 87 35 45 0.00105685 0.000666853 0.000796853 0.331352 0.00118345 0.000475253 0.000711318 0.523537 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 86 33 46 0.000997102 0.000709494 0.000805363 0.312391 0.00117978 0.000484607 0.00071633 0.51874 86 33 46 0.000997102 0.000709494 0.000805363 0.312391 0.00117978 0.000484607 0.00071633 0.51874 86 32 46 0.00144502 0.000577058 0.00086638 0.524286 0.00118822 0.000480466 0.000716384 0.525261 86 32 46 0.00144502 0.000577058 0.00086638 0.524286 0.00118822 0.000480466 0.000716384 0.525261 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 29 45 0.00159702 0.000373003 0.00078101 0.732022 0.00120427 0.000464151 0.000710859 0.54041 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 |
| Line 198: | Line 239: |
Back to list of all tutorials | Back to course page | Previous
Tract statistics
Please bear with us while we update the tutorial data set for the new version of TRACULA. The new tutorial goes beyond what was possible with the tutorial data set that was previously available for download. For now, please ignore any references to the tutorial data and try to follow along using your own data. Thank you for your patience. |
Remember...
For each new terminal that you open, you must do:
export SUBJECTS_DIR=$TUTORIAL_DATA/diffusion_tutorial/fs cd $TUTORIAL_DATA/diffusion_tutorial
This section of the tutorial will teach you how to extract statistics on anisotropy and diffusivity measures for the white-matter pathways reconstructed by TRACULA. There are two types of statistics files that are created for each white-matter pathway:
pathstats.overall.txt - This file contains diffusion measures averaged over the whole pathway.
pathstats.byvoxel.txt - This file contains diffusion measures averaged at consecutive cross-sections of the pathway, resulting in an along-tract profile of each diffusion measure. This type of analysis is referred to as pointwise assessment of streamline tractography attributes (PASTA).
These are plain text files that you can open with a text editor, such as gedit on Linux, or with the command open -e on a Mac. They are saved in the TRACULA output directory of each individual tract.
Anisotropy and diffusivity averaged over an entire pathway
Examine the whole-tract statistics for the left inferior longitudinal fasciculus (ILF) of subject1, by runnning:
cat trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt
This file will look like this:
# Title Pathway Statistics # # generating_program /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats # cvs_version ayendiki-local # cmdline /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats --intrc /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr --dtbase /space/erebus/1/users/data/trc/subject1/dmri/dtifit --path lh.ilf --subj subject1 --invox path.ref.txt --out /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt --outvox /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.byvoxel.txt # sysname Linux # hostname compute-0-124.nmr.mgh.harvard.edu # machine x86_64 # user ayendiki # anatomy_type pathway # # subjectname subject # pathwayname lh.ilf # Count 1500 Volume 1996 Len_Min 53 Len_Max 129 Len_Avg 79.9787 Len_Center 89 AD_Avg 0.00120044 AD_Avg_Weight 0.00121209 AD_Avg_Center 0.00123318 RD_Avg 0.000467543 RD_Avg_Weight 0.000458132 RD_Avg_Center 0.000549716 MD_Avg 0.000711842 MD_Avg_Weight 0.000709453 MD_Avg_Center 0.000777538 FA_Avg 0.540167 FA_Avg_Weight 0.553683 FA_Avg_Center 0.477498
This text file contains various diffusion measures, averaged over the entire pathway (in this example, the left ILF). The measures include:
- Number of sample paths drawn from the probability distribution of the pathway (Count)
- Volume of the probability distribution of the pathway (in voxels)
- Maximum, minimum, and average length of the sample paths (in voxels)
- Length of the highest-probability (a.k.a. maximum a posteriori) path
- Axial diffusivity (average over the entire support of the path distribution, weighted average over the entire support of the path distribution, and average over highest-probability path only)
- Radial diffusivity (as above)
- Mean diffusivity (as above)
- Fractional anisotropy (as above)
Concatenate for group analyses
Measures can be extracted from these files and concatenated across subjects, e.g., for group analysis. Specifically, the text files can be converted into a table using the command tractstats2table and then used for doing GLM analyses with mri_glmfit or any other statistical software (SPSS, Excel, Statview etc.)
Before using tractstats2table, you have to create a text file that lists the full path to the pathstats.overall.txt files from all subjects that you want to include in your analysis. For example, if you wanted to analyze whole-tract measures from the left ILF of 3 subjects, you would create a text file that looks like the following:
trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt trc/subject2/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt trc/subject3/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt
Create a text file called lh.ilf.list using the command below. Copy and paste the list above into this file, then save the file and exit the editor.
gedit lh.ilf.list &
ON MACS, RUN:
open -e lh.ilf.list &
The following command will use this list to create a table with the diffusion measures from all the subjects listed above:
tractstats2table --load-pathstats-from-file lh.ilf.list --overall --tablefile lh.ilf.All.table
The argument to the --load-pathstats-from-file option specifies the text file that contains the list of all the statistics files that will be combined.
Take a look at the resulting table file:
gedit lh.ilf.All.table &
ON MACS, RUN:
open -e h.ilf.All.table &
Instead of extracting all measures included in the pathstats.overall.txt files, we are usually interested only in a few specific measures. For example, to extract only the average FA for each subject, do the following:
tractstats2table --load-pathstats-from-file $lh.ilf.list --overall --only-measures FA_Avg --tablefile lh.ilf.FA_Avg.table
The argument to the --only-measures option specifies which measure we want to extract from the statistics files. Instead of FA_Avg, this could be the name of any of the measures included in pathstats.overall.txt.
OPTIONAL: You can look at these tables in OpenOffice (or any other spreadsheet program). For example, to open the file lh.ilf.All.table in OpenOffice, do the following :
localc lh.ilf.All.table
You can then use these stats tables as input to mri_glmfit, to run perform group analysis.
Anisotropy and diffusivity along the trajectory of a pathway
Now examine the along-tract (PASTA) statistics for the left ILF, by running:
cat trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.byvoxel.txt
# Title Pathway Statistics # # generating_program /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats # cvs_version ayendiki-local # cmdline /usr/local/freesurfer/7.2.0-beta/bin/dmri_pathstats --intrc /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr --dtbase /space/erebus/1/users/data/trc/subject1/dmri/dtifit --path lh.ilf --subj subject1 --invox path.ref.txt --out /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.overall.txt --outvox /space/erebus/1/users/data/trc/subject1/dpath/lh.ilf_avg16_syn_bbr/pathstats.byvoxel.txt # sysname Linux # hostname compute-0-124.nmr.mgh.harvard.edu # machine x86_64 # user ayendiki # anatomy_type pathway # # subjectname subject1 # pathwayname lh.ilf # # pathway start x y z AD RD MD FA AD_Avg RD_Avg MD_Avg FA_Avg 97 78 16 0.000867977 0.000725794 0.000773188 0.108892 0.00107168 0.000599572 0.000756942 0.363722 97 78 16 0.000867977 0.000725794 0.000773188 0.108892 0.00107168 0.000599572 0.000756942 0.363722 97 77 16 0.000832794 0.000712196 0.000752395 0.098711 0.00106892 0.000599915 0.000756251 0.363217 97 77 16 0.000832794 0.000712196 0.000752395 0.098711 0.00106892 0.000599915 0.000756251 0.363217 96 77 17 0.000778157 0.000713259 0.000734892 0.0524794 0.00106822 0.000599456 0.000755712 0.364947 96 76 17 0.000872096 0.000706771 0.000761879 0.136309 0.00106737 0.00059909 0.000755185 0.365999 96 76 17 0.000872096 0.000706771 0.000761879 0.136309 0.00106737 0.00059909 0.000755185 0.365999 96 75 18 0.00106175 0.000671836 0.000801806 0.326543 0.00106621 0.000593117 0.000750814 0.370047 96 74 19 0.00084758 0.000742526 0.000777544 0.146359 0.00107689 0.000564038 0.000734991 0.399445 96 74 19 0.00084758 0.000742526 0.000777544 0.146359 0.00107689 0.000564038 0.000734991 0.399445 96 73 20 0.00105676 0.000582545 0.000740616 0.379989 0.00109462 0.000529155 0.000717645 0.442998 96 73 20 0.00105676 0.000582545 0.000740616 0.379989 0.00109462 0.000529155 0.000717645 0.442998 96 73 21 0.00106829 0.000535296 0.000712959 0.446277 0.00110578 0.000510944 0.000709221 0.466119 96 73 21 0.00106829 0.000535296 0.000712959 0.446277 0.00110578 0.000510944 0.000709221 0.466119 96 72 21 0.00112798 0.000524413 0.000725602 0.490875 0.00109594 0.000500484 0.000698966 0.476156 96 72 22 0.00100371 0.000518413 0.00068018 0.466196 0.00109198 0.000491746 0.000691821 0.484992 96 71 22 0.00110846 0.000516744 0.000713983 0.496827 0.00107691 0.000488262 0.000684476 0.487281 96 70 23 0.000963374 0.000468995 0.000633788 0.515347 0.00103651 0.000500421 0.000679117 0.471307 96 70 24 0.000952905 0.00052878 0.000670155 0.464126 0.00102183 0.000507845 0.000679174 0.463955 96 69 25 0.000941334 0.000510755 0.000654282 0.452879 0.0010153 0.000520488 0.000685426 0.451515 96 69 25 0.000941334 0.000510755 0.000654282 0.452879 0.0010153 0.000520488 0.000685426 0.451515 96 68 25 0.000937902 0.000498823 0.000645183 0.473998 0.0010151 0.000521116 0.000685777 0.45532 96 67 26 0.00114779 0.000422178 0.00066405 0.597208 0.00106285 0.000495767 0.000684795 0.488664 96 67 26 0.00114779 0.000422178 0.00066405 0.597208 0.00106285 0.000495767 0.000684795 0.488664 96 66 27 0.00119928 0.000456432 0.000704049 0.567628 0.00113942 0.000464739 0.000689633 0.535918 96 66 27 0.00119928 0.000456432 0.000704049 0.567628 0.00113942 0.000464739 0.000689633 0.535918 96 65 27 0.00141769 0.000388493 0.000731558 0.691121 0.00118206 0.000446907 0.000691963 0.56636 96 64 28 0.00139466 0.000325184 0.000681676 0.734954 0.00123269 0.000410431 0.000684517 0.612061 96 64 28 0.00139466 0.000325184 0.000681676 0.734954 0.00123269 0.000410431 0.000684517 0.612061 96 63 28 0.00143303 0.000312452 0.000685979 0.752811 0.00126731 0.000393664 0.000684881 0.636549 95 62 29 0.00130884 0.000290893 0.000630209 0.745983 0.00129524 0.000387733 0.000690235 0.648474 94 62 29 0.00164671 0.00076911 0.00106164 0.446289 0.00129674 0.000390088 0.000692304 0.647809 94 61 29 0.00172456 0.000745308 0.00107172 0.485371 0.00127988 0.000398742 0.000692451 0.638261 94 60 30 0.00140745 0.000880179 0.00105594 0.281852 0.00125246 0.000428793 0.00070335 0.606134 94 60 30 0.00140745 0.000880179 0.00105594 0.281852 0.00125246 0.000428793 0.00070335 0.606134 94 59 31 0.00155751 0.000862578 0.00109422 0.356395 0.00121421 0.000460178 0.000711525 0.56957 94 58 32 0.00130825 0.000701944 0.000904047 0.376433 0.00120559 0.000462724 0.00071035 0.562654 94 58 32 0.00130825 0.000701944 0.000904047 0.376433 0.00120559 0.000462724 0.00071035 0.562654 94 57 33 0.00127009 0.000453103 0.000725431 0.580671 0.001259 0.000436023 0.000710351 0.597881 94 56 34 0.00128442 0.000408392 0.0007004 0.624911 0.00131045 0.000409303 0.000709684 0.635395 94 56 34 0.00128442 0.000408392 0.0007004 0.624911 0.00131045 0.000409303 0.000709684 0.635395 94 55 34 0.00129227 0.00046328 0.00073961 0.577816 0.0013183 0.000405609 0.000709842 0.640691 94 54 35 0.0012703 0.000405959 0.000694073 0.624761 0.00132773 0.000405781 0.000713098 0.639696 94 54 36 0.0012825 0.000416241 0.000704992 0.62111 0.00131601 0.00041376 0.000714509 0.62888 94 54 36 0.0012825 0.000416241 0.000704992 0.62111 0.00131601 0.00041376 0.000714509 0.62888 93 53 36 0.00139747 0.000604594 0.000868887 0.488666 0.00130335 0.000422725 0.000716265 0.616271 93 52 36 0.00135543 0.000585794 0.000842337 0.488772 0.0012934 0.000424784 0.000714323 0.612202 93 52 37 0.00123236 0.000470209 0.000724259 0.552723 0.00128567 0.000433751 0.000717722 0.600983 93 51 38 0.00114102 0.000441359 0.00067458 0.550956 0.00128498 0.000448106 0.000727063 0.587959 93 50 38 0.00115213 0.00050062 0.000717791 0.500001 0.00128811 0.00045668 0.00073382 0.580909 93 50 39 0.00106929 0.000415801 0.000633631 0.546817 0.00129122 0.00045262 0.000732149 0.585613 92 49 39 0.00139456 0.000653969 0.000900833 0.448212 0.00129614 0.000456588 0.000736437 0.583877 92 48 39 0.00121658 0.000614545 0.000815223 0.410542 0.00129869 0.00045225 0.000734397 0.588016 92 48 39 0.00121658 0.000614545 0.000815223 0.410542 0.00129869 0.00045225 0.000734397 0.588016 92 47 40 0.00118115 0.000483847 0.000716281 0.521239 0.0013064 0.00044644 0.000733091 0.593569 91 46 41 0.00132381 0.000679848 0.000894501 0.400171 0.00130968 0.000445202 0.000733361 0.593125 91 46 41 0.00132381 0.000679848 0.000894501 0.400171 0.00130968 0.000445202 0.000733361 0.593125 91 45 41 0.00113019 0.000529645 0.000729826 0.454707 0.001325 0.000431276 0.000729183 0.607594 91 44 41 0.00117487 0.000464237 0.000701113 0.530762 0.00134519 0.00041734 0.000726622 0.623841 90 43 41 0.00121382 0.000492301 0.000732807 0.519596 0.00135435 0.00040553 0.000721804 0.635504 90 42 42 0.00141832 0.000419311 0.000752312 0.650028 0.00135835 0.000396281 0.000716967 0.644296 90 41 43 0.00149731 0.000356095 0.0007365 0.72266 0.00135424 0.000393535 0.00071377 0.64191 89 40 43 0.001381 0.000513018 0.000802344 0.564421 0.00132657 0.000407598 0.000713924 0.624407 89 40 43 0.001381 0.000513018 0.000802344 0.564421 0.00132657 0.000407598 0.000713924 0.624407 90 39 43 0.00157585 0.000295255 0.000722119 0.787805 0.00130086 0.000413205 0.00070909 0.613173 89 38 43 0.00138324 0.000383207 0.000716551 0.674917 0.00125725 0.000434189 0.000708544 0.581023 89 38 44 0.00124692 0.000510601 0.000756042 0.514663 0.001253 0.000438098 0.000709733 0.575357 89 37 45 0.00118923 0.00056238 0.000771331 0.44532 0.00122624 0.00044834 0.000707639 0.557656 88 37 45 0.00109356 0.000633894 0.000787115 0.365015 0.00123292 0.000451307 0.000711844 0.55797 88 36 45 0.00103814 0.000676695 0.000797177 0.316449 0.00120901 0.000458568 0.000708718 0.544602 88 35 45 0.00114301 0.000589147 0.000773767 0.391688 0.00118172 0.000467419 0.000705519 0.529077 87 35 45 0.00105685 0.000666853 0.000796853 0.331352 0.00118345 0.000475253 0.000711318 0.523537 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 87 34 45 0.00125167 0.000637577 0.000842276 0.399288 0.001166 0.000479059 0.000708037 0.516315 86 33 46 0.000997102 0.000709494 0.000805363 0.312391 0.00117978 0.000484607 0.00071633 0.51874 86 33 46 0.000997102 0.000709494 0.000805363 0.312391 0.00117978 0.000484607 0.00071633 0.51874 86 32 46 0.00144502 0.000577058 0.00086638 0.524286 0.00118822 0.000480466 0.000716384 0.525261 86 32 46 0.00144502 0.000577058 0.00086638 0.524286 0.00118822 0.000480466 0.000716384 0.525261 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 31 46 0.00145415 0.000575363 0.000868292 0.528119 0.00121054 0.000477254 0.000721682 0.534651 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 30 46 0.00161654 0.000434083 0.000828234 0.688725 0.00121171 0.000469266 0.000716745 0.540934 85 29 45 0.00159702 0.000373003 0.00078101 0.732022 0.00120427 0.000464151 0.000710859 0.54041 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 84 29 45 0.00149992 0.000433867 0.000789218 0.665445 0.0012176 0.000470298 0.000719398 0.53854 # pathway end
This text file contains various diffusion measures, one row for each position along the trajectory of the path. The first three entries in each row are the x, y, z coordinates in native diffusion space. The next four entries are the axial diffusivity, radial diffusivity, mean diffusivity, and fractional anisotropy at that position on the maximum a posteriori path. The last four entries are the axial diffusivity, radial diffusivity, mean diffusivity, and fractional anisotropy at the same position, averaged over all sampled paths.
Points are ordered according to the following convention (see also along-tract plots):
- For pathways with a right-left orientation:
- If the pathway is inter-hemispheric: From right to left
- If the pathway is intra-hemispheric: From medial to lateral
- For pathways with an anterior-posterior orientation: From anterior to posterior
- For pathways with a superior-inferior orientation: From superior to inferior
Converting pathstats.byvoxel.txt files to a table for group analyses
You can combine the pathstats.byvoxel.txt files from multiple subjects, to use as input for along-tract group analysis, by running the following:
trac-all -stat -c $TUTORIAL_DATA/diffusion_tutorial/dmrirc.tutorial
This will create a directory named stats under the main TRACULA output directory and save one table per tract per diffusion measure. In these tables, each row is a different position along the trajectory of the tract and each column is a different subject.
To examine the table of average FA along the left ILF, do the following:
gedit trc/stats/lh.ilf.avg16_syn_bbr.FA_Avg.txt &
ON MACS, RUN:
open -e trc/stats/lh.ilf.avg16_syn_bbr.FA_Avg.txt &
The contents of this file will look like this:
elmo.2005 elmo.2008 elmo.2012 NaN NaN 0.378069 NaN NaN 0.378492 0.41336 NaN 0.388788 0.429444 0.377014 0.4164 0.444207 0.392511 0.438363 0.452544 0.394491 0.445661 0.449954 0.388662 0.435143 0.447727 0.407221 0.426532 0.432298 0.428718 0.421008 0.419581 0.425808 0.403944 0.473702 0.40381 0.371384 0.494473 0.387531 0.353198 0.4767 0.414657 0.366791 0.506063 0.453886 0.412596 0.583493 0.519466 0.483278 0.624087 0.57846 0.549247 0.647596 0.600432 0.569196 ...
The first row tells you the subject names for the corresponding columns of FA values.
You can use these stats tables to run a group analysis on the tracts.
Note: In addition to the above tables, the stats directory will also contain a log file for each pathway. You can examine the log file, which contains the full output of the trac-all -stat command, to see if any pathways were flagged as outliers. If this happens, there will be a line in the log file with "Found outlier path:" and the name of the subject. These are paths that were found to differ excessively from those of the other subjects. You can check this information to find out if the reconstruction of this tract failed for any subjects.
For example, you can examine the log file for the left ILF by doing:
gedit trc/stats/lh.ilf.avg16_syn_bbr.log &
ON MACS, RUN:
open -e trc/stats/lh.ilf.avg16_syn_bbr.log &
Visualizing results from statistical analyses along each pathway
By default, TRACULA uses the mean of its manually annotated training streamlines to determine the positions along each of the 42 pathways where diffusion measures will be projected. These mean streamlines are saved in template space (the template that is the target of the inter-subject registration) and mapped to each individual subject after the individual subjects pathways are reconstructed, to extract the along-tract measures. Note that these mean paths are only used to determine how each pathway will sliced up into into cross-sections where along-tract measures will be averaged, and to ensure an equal number of cross-sections for all subjects.
To view all 42 mean paths in template space, run the following:
freeview -v $FREESURFER_HOME/trctrain/hcp/MGH35_HCP_FA_template.nii.gz \
-t $FREESURFER_HOME/trctrain/hcp/syn/*.mean.trkYou should see something like this in the 3D view:
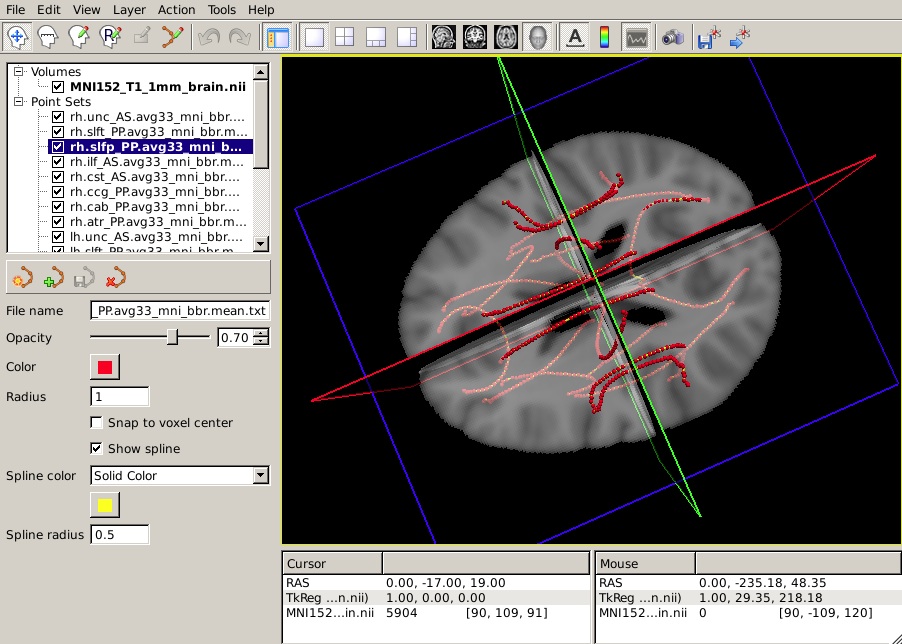
The following information is based on hypothetical statistical data. The text file of p-values is not available so do not expect your end result to look identical to this image.
Let's say that you have performed a statistical analysis on your subjects in your statistical software of choice. If you save the p-values corresponding to each position along a tract to a simple text file, you can now display these p-values as a heat map on the corresponding mean path. (It is assumed that the number of p-values in your text file is equal to the number of points on the mean path, which is the same as the number of rows of values in the group tables produced by trac-all -stat. If not, display may be problematic.)
To display the p-values from your hypothetical analysis on one of the mean paths, do the following:
Select the mean path of your choice on the panel in the top left of the freeview window. On the left menu, click on Show spline.
From the Spline color menu, choose Heatscale.
From the Scalar map menu, choose Load... and select the text file that contains the p-values from your statistical analysis.
To hide the waypoints, which show up as small spheres, increase the Spline radius so that it is greater than the Radius of the waypoints. (In the example below, we have set the Spline radius to 2, while the Radius of the waypoints is 1.)
Set the Min, Mid, and Max of the heatscale to threshold the p-values as you wish.
You can do this for as many of the tracts as you have performed statistical analyses on. The end result will look something like this in the 3D view:
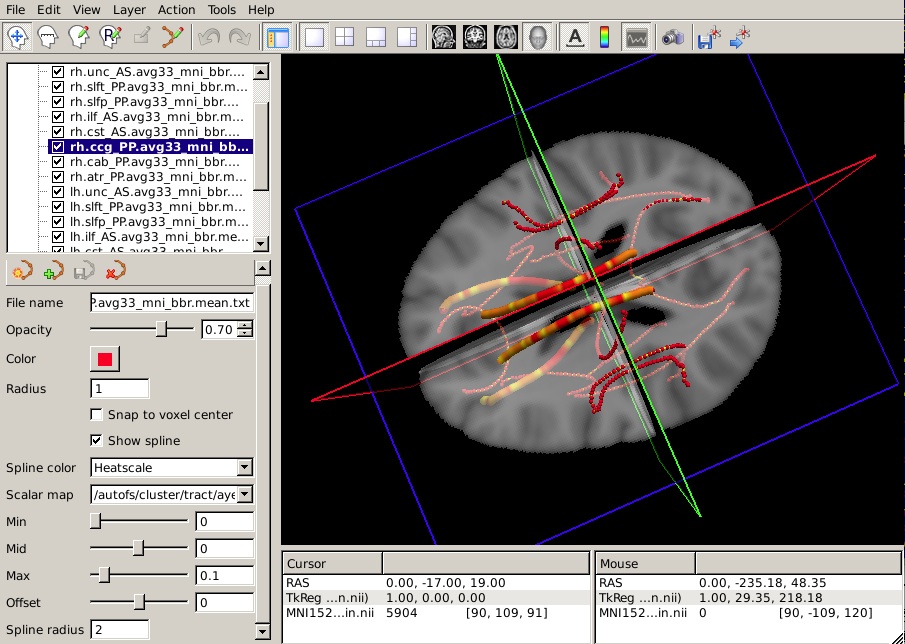
Summary
By the end of this page, you should know how to:
- Extract statistics (anisotropy and diffusivity measures), either averaged over an entire white-matter pathway, or as a function of position along the trajectory of the pathway
- Combine statistics into tables for group analysis
- Perform whole-tract and along-tract statistical analysis
- Visualize the outputs of along-tract statistical analysis in freeview
Quiz
You can test your knowledge of this tutorial by clicking here for a quiz!
Back to list of all tutorials | Back to course page | Previous
