|
Size: 7552
Comment:
|
← Revision 39 as of 2026-04-15 10:48:13 ⇥
Size: 8827
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| <<BR>> <<BR>> '''''Important: please download the latest development version of FreeSurfer to use this package''''' |
|
| Line 14: | Line 17: |
| A preprint of the manuscript describing the atlas and segmentation method used in this module can be found [[https://www.biorxiv.org/content/10.1101/2024.02.05.579016v1|on bioRxiv.]] | Relevant publications: |
| Line 16: | Line 19: |
| "A probabilistic histological atlas of the human brain for MRI segmentation", Casamitjana et al., Nature, 2025. [[https://www.nature.com/articles/s41586-025-09708-2|Paper on nature.com.]] <<BR>> "Fast segmentation with the NextBrain histological atlas", Puonti et al., Imaging Neuroscience, 2026 (accepted). [[https://www.biorxiv.org/content/10.1101/2025.09.22.673638v1|Preprint available here.]] <<BR>> <<BR>> |
|
| Line 19: | Line 30: |
| 3. Usage ('full' Bayesian version, '''not recommended''') 4. Usage (faster versions) 5. Frequently asked questions (FAQ) |
3. Basic usage 4. Outputs 5. Advanced options 6. Frequently asked questions (FAQ) |
| Line 25: | Line 37: |
| This module uses our new probabilistic atlas of the human brain to segment 333 distinct ROIs per hemisphere on in vivo scans. Segmentation relies on a Bayesian algorithm and is thus robust against changes in MRIi pulse sequence (e.g., T1-weighted, T2-weighted, FLAIR, etc). Sample slices of the atlas and the segmentation of the sample subject "bert" are shown below: | This module uses NextBrain, our new probabilistic atlas of the human brain, to segment ~300 distinct ROIs per hemisphere on in vivo or ex vivo scans (including single hemispheres). Segmentation relies on a Bayesian algorithm and is thus robust against changes in MRI pulse sequence (e.g., T1-weighted, T2-weighted, FLAIR, etc). Sample slices of the atlas and the segmentation of the sample subject "bert" are shown below: |
| Line 29: | Line 41: |
| The first time you run this module, it will prompt you to download the atlas. Follow the instructions on the screen to obtain the files. | The first time you run this module, it will prompt you to download the atlas. Follow the instructions on the screen to obtain the atlas files. In addition: this module calls [[https://surfer.nmr.mgh.harvard.edu/fswiki/SuperSynth|mri_super_synth]]; if you have never used this command before, it will also prompt you to download a model file. |
| Line 32: | Line 46: |
| === 3. Usage ('full' Bayesian version; not recommended as it is very slow) === To segment a brain MRI scan, |
=== 3. Basic usage === The entry point of the module is the command '''mri_histo_atlas_segment_fireants''', which implements [[https://www.biorxiv.org/content/10.1101/2025.09.22.673638v1|the fast version of the algorithm]]. This version relies on [[https://arxiv.org/abs/2404.01249|FireANTs (Jena et al)]] for fast nonlinear registration of the atlas. The command line is: |
| Line 35: | Line 51: |
| mri_histo_atlas_segment INPUT_SCAN OUTPUT_DIRECTORY ATLAS_MODE GPU THREADS [BF_MODE] [GMM_MODE] | mri_histo_atlas_segment_fireants --i INPUT_SCAN --o OUTPUT_DIRECTORY --device [cpu/cuda] --side [left/right] --mode [invivo/cerebrum/hemi/exvivo] |
| Line 37: | Line 53: |
| where: | |
| Line 39: | Line 54: |
| ''INPUT_SCAN'': scan to process, in mgz or nii(.gz) format. | * ''INPUT_SCAN'': scan to process, in mgz or nii(.gz) format. |
| Line 41: | Line 56: |
| ''OUTPUT_DIRECTORY'': directory where segmentations, volume files, etc, will be created (more on this below). | * ''OUTPUT_DIRECTORY'': directory where segmentations, volume files, etc, will be created (more on this below). |
| Line 43: | Line 58: |
| ''ATLAS_MODE'' : full (atlas with all 333 labels) or simplified (simpler brainstem protocol; recommended) | * ''DEVICE'': set to cpu or cuda. |
| Line 45: | Line 60: |
| ''GPU'': set to 1 to use the GPU ('''we highly recommend using a GPU to run this module; without a GPU, running this module on a single scan can take tens for hours'''). The GPU requirements depend on the image but are about 24GB of memory. | * ''SIDE'': left or right. If you're analyzing both sides, you're better off running them sequentially (rather than in parallel) since the SuperSynth preprocessing will be reused when processing the second hemisphere. |
| Line 47: | Line 62: |
| ''THREADS'': number of CPU threads used by the code (set to -1 to use all available threads). ''BF_MODE'' (optional): bias field mode: dct (default), polynomial, or hybrid. ''GMM_MODE'' (optional): gaussian mixture model (GMM) model must be 1mm unless you define your own (see FAQ below). |
* ''MODE'': type of scan: invivo, exvivo, cerebrum (ex vivo without brainstem or cerebellum), hemi (ex vivo with single cerebral hemisphere). |
| Line 54: | Line 65: |
| === 4. Outputs === | |
| Line 55: | Line 67: |
| For example, you can segment bert with a GPU and 8 CPU threads with: | The output directory will contain the following files: |
| Line 57: | Line 69: |
| {{{ mri_histo_atlas_segment $SUBJECTS_DIR/bert/mri/orig.mgz $SUBJECTS_DIR/bert/mri/histo_atlas_segmentation/ simplified 1 8 }}} |
* ''seg.[left/right].nii.gz'': segmentation of left/right hemisphere |
| Line 61: | Line 71: |
| In the output directory, you will find: | * ''lut.txt'': the lookup table to visualize seg.[left/right].nii.gz, for convenience |
| Line 63: | Line 73: |
| ''bf_corrected.mgz'': a bias field corrected version of the input scan. | * ''vols.[left/right].csv'': files with volumes of the brain regions segmented by the atlas, in CSV format. |
| Line 65: | Line 75: |
| ''SynthSeg.mgz'': [[https://surfer.nmr.mgh.harvard.edu/fswiki/SynthSeg|SynthSeg]] segmentation of the input (which we use in preprocessing and to initialize Gaussian parameters). | * ''SuperSynth'': directory with segmentation of the scan at the whole structure level. |
| Line 67: | Line 77: |
| ''MNI_registration.mgz'': [[https://surfer.nmr.mgh.harvard.edu/fswiki/EasyReg|EasyReg]] registration to MNI space, use in preprocessing. | Additional flags: if advanced options are used (more details below). <<BR>> |
| Line 69: | Line 80: |
| ''seg_[left/right].mgz'': segmentation into 333 ROIs of the left and right hemisphere, respectively. | |
| Line 71: | Line 81: |
| ''vols_[left/right].csv'': CSV spreadsheet with the volumes of the different ROIs, computed from the posteriors (soft segmentations). | === 5. Advanced options === |
| Line 73: | Line 83: |
| ''lookup_table.txt'': FreeSurfer lookup table mapping label indices to brain anatomy. You need it when visualizing the segmentations with Freeview. | The code also accepts the following optional flags: * --bf_mode: Decides the bias field basis function model. Options: dct (default), polynomial, hybrid. * --write_rgb: Save an RGB image based on the posterior probabilites to disk. * --write_bias_corrected: Save the bias corrected input image to disk. * --device_registration: Define a different device for the registration. Can be used to save GPU memory when working with an GPU with limited memory. Options: cpu, cuda. Default is the same as --device. * --threads: Control the number of cpu thread used to run the algorithm. Default value is -1, which uses all available threads. * --skip: An integer skipping (downsampling) factor for estimating the model parameters. More skipping saves memory, but sacrifices accuracy. Default: 1 (no skipping). * --resolution: The resolution of the output segmentation. By default 0.4mm, which is higher than the typical input scan, to reduce aliasing. * --smoothing_steps_HRmask: Number of smoothing steps used when upsampling the 1mm brain mask from BrainFM. More smoothing makes the outer border less jagged, but too much smoothing reduces accuracy. Default: 3. * --skip_bf: Skip the bias field correction. Can be used to save memory if the input scan is already bias corrected or does not have a bias field (non MRI modality). * --smooth_grad_sigma: Gradient field smoothing parameter for the nonlinear FireAnts registration. Default: 1.0. * --smooth_warp_sigma: Warp field smoothing parameter for the nonlinear FireAnts registration. Default: 0.25. * --optimizer_lr: Learning rate for the nonlinear FireAnts registration optimizer. Default: 0.5. * --cc_kernel_size: Size of the window for calculating the cross-correlation registration metric. Default: 7. * --rel_weight_labeldiff: Relative weight for the Dice loss metric in the nonlinear registration. Default: 2.5. * --save_atlas_nonlinear_reg: Save the nonlinearly registered atlas. Default: false. * --save_field: Save the nonlinear deformation field. Default: false. * --save_jacobian: Save the Jacobian determinant (log10) of the deformation field. Default: false. * --yaml_path: path of custom YAML files to define groups of ROIs Some notes: * If you are running out of memory, using --skip 2 can help without sacrificing much accuracy. * The defaults --smooth_grad_sigma 1 and --smooth_warp_sigma 0.25 are pretty liberal and can cope with massive deformation, e.g., as in the Hip-CT images shown in the paper "Fast segmentation with the NextBrain". If you are working with a population without very strong atrophy or deformation, you can multiply those values by 2 in order to get more regular atlas deformation fields (you can explore the deformation with the --save_jacobian option). Also: you can flexibly change the groupings of the modeled structures using the .yaml files under the /data_simplified folder. The structure groupings for the Gaussian Mixture modeled are controlled by two files: gmm_components_fireants.yaml and combined_atlas_labels_fireants.yaml. Let's say, as an example, that you wanted to add the internal segment of globus pallidus (label 206) as its own structure. To model it separately, you would first create a new class, called e.g., Internal Segment Pallidum, in the combined_atlas_labels_fireants.yaml file, and list label 206 under that structure (while removing it from the pallidum class). Next, you would add the class, with exactly the same name, to the gmm_components_fireants.yaml file and decide how many Gaussian distributions should be used to model its intensities. To make the non-linear registration aware of the contrast, you would add the structure, again with exactly the same name, to the file called recipe_intensities_cheating_image_fireants.yaml, and decide how its intensity should be generated from the seven structures than can be always reliably segmentation using BrainFM (see the file for examples). |
| Line 77: | Line 114: |
| === 4. Usage (faster versions) === | === 6. Frequently asked questions (FAQ) === |
| Line 79: | Line 116: |
| '''We just released these, so please bear with us (and definitely let us know!) if the code is buggy or crashes.''' | * '''I have an ex vivo hemisphere with cerebellum and/or brainstem''' |
| Line 81: | Line 118: |
| We also distribute two faster version, where the atlas deformation is pre-computed and then kept constant during the optimization, such that we only need to run the EM algorithm once for the Gaussian parameters and that is it. |
If you use the hemi mode, you will not get the cerebellum or brainstem. Use the exvivo mode instead (with the caveat that you may lose some voxels around the medial wall, which may get assigned to the contralateral hemisphere). |
| Line 85: | Line 120: |
| There are two sub-versions. One uses [[https://arxiv.org/abs/2404.01249|FireANTs (Jena et al)]] to register to the target scan a synthetic cartoon derived from the atlas. This version actively tries to fit smaller structures and subregions. The command line is: |
* '''Can the exvivo model handle arbitrary orientations of the input''' |
| Line 89: | Line 122: |
| {{{ mri_histo_atlas_segment_fireants INPUT_SCAN OUTPUT_DIRECTORY GPU THREADS [BF_MODE] }}} |
No, it cannot. You need to manually reorient the brain to RAS (e.g., with Freeview). |
| Line 93: | Line 124: |
| The other version uses [[https://arxiv.org/abs/2404.01249|SynthMorph (Hoffmann et al., Imaging Neuroscience, 2024)]], a neural network for image registration. This version is faster and does OK for 1mm isotropic scans. However, SynthMorph relies heavily on fitting the boundaries of whole structures and does not the map smaller regions as well (e.g., thalamic nuclei). Therefore, we do not recommend it for images with resolution better than 1mm isotropic (e.g., ex vivo scans). |
* '''Do I need a GPU?''' |
| Line 100: | Line 126: |
| {{{ mri_histo_atlas_segment_fast INPUT_SCAN OUTPUT_DIRECTORY GPU THREADS [BF_MODE] }}} |
Certainly not! The code should run in less than half an hour on any semi-modern workstation, if you allocate enough threads (or about two hours for an ex vivo scan at 0.25mm resolution). |
| Line 104: | Line 128: |
| For both commands, the options are similar to mri_histo_atlas_segment, but the atlas and gmm modes are always 'simplified' and '1mm', respectively. The output files in the output directory follow the same convention. |
* '''What happened to the "full Bayesian" and "SynthMorph" versions?''' |
| Line 107: | Line 130: |
| These faster versions are particularly useful if you are running the code on the CPU rather than CPU. On a semi-modern desktop, the run time should be less than an hour; note that mri_histo_atlas_segment_fireants runs once per hemisphere, whhile mri_histo_atlas_segment_fast segments both hemispheres in a single run. |
To simplify the codebase, we are focusing on this method, which is fast but also versatile in terms of modeling / registration (as opposed to SynthMorph). <<BR>> <<BR>> |
| Line 115: | Line 139: |
=== 5. Frequently asked questions (FAQ) === * '''Do I really need a GPU for the 'full' Bayesian version?''' Technically, no. In practice, yes. On a modern GPU, the code runs in an hour or less. On the CPU, it depends on the number of threads, but it can easily take a whole day or more. * '''Do I need a GPU for the faster versions?''' Certainly no! The code should run in less than an hour on any semi-modern workstation, if you allocate enough threads (or about two hours for an ex vivo scan at 0.25mm resolution). * '''What do the BF_MODE and GMM_MODE arguments do in the full Bayesian version?''' You should not need to touch these, but the BF_MODE changes the set of basis functions for bias field correction and you could potentially try tinkering with it if the bias field correction fails (i.e., if bf_corrected.mgz has noticeable bias). GMM_MODE allows you to change the grouping of ROIs into tissue classes (advanced mode!). The GMM model is crucial as it determines how different brain regions are grouped into tissue types for the purpose of image intensity modeling. This is specified though a set of files that should be found under 'data' in the atlas directory: ''data/gmm_components_[GMM_MODE].yaml'': defines tissue classes and specificies the number of components of the corresponding GMM ''data/combined_aseg_labels_[GMM_MODE].yaml'' defines the labels that belong to each tissue class ''data/combined_atlas_labels_[GMM_MODE].yaml'' defines FreeSurfer ('aseg') labels that are used to initialize the parameters of each class. Note that, in in dev versions newer than August 23 2024, these files are located under data_full and data_simplified (one directory per atlas / protocol). We distribute a GMM_MODE named "1mm" that we have used in our experiments, and which is the default mode of the code. If you want to use your own model, you will need to create another triplet of files of your own (use the 1mm version as template). <<BR>> |
Bayesian Segmentation with Histological Atlas "NextBrain"
Visit the homepage of the NextBrain project for further information on this atlas.
Important: please download the latest development version of FreeSurfer to use this package
Author: Juan Eugenio Iglesias
E-mail: jiglesiasgonzalez [at] mgh.harvard.edu
Rather than directly contacting the author, please post your questions on this module to the FreeSurfer mailing list at freesurfer [at] nmr.mgh.harvard.edu
Relevant publications:
"A probabilistic histological atlas of the human brain for MRI segmentation", Casamitjana et al., Nature, 2025. Paper on nature.com.
"Fast segmentation with the NextBrain histological atlas", Puonti et al., Imaging Neuroscience, 2026 (accepted). Preprint available here.
Contents
- General Description
- Installation
- Basic usage
- Outputs
- Advanced options
- Frequently asked questions (FAQ)
1. General Description
This module uses NextBrain, our new probabilistic atlas of the human brain, to segment ~300 distinct ROIs per hemisphere on in vivo or ex vivo scans (including single hemispheres). Segmentation relies on a Bayesian algorithm and is thus robust against changes in MRI pulse sequence (e.g., T1-weighted, T2-weighted, FLAIR, etc). Sample slices of the atlas and the segmentation of the sample subject "bert" are shown below:
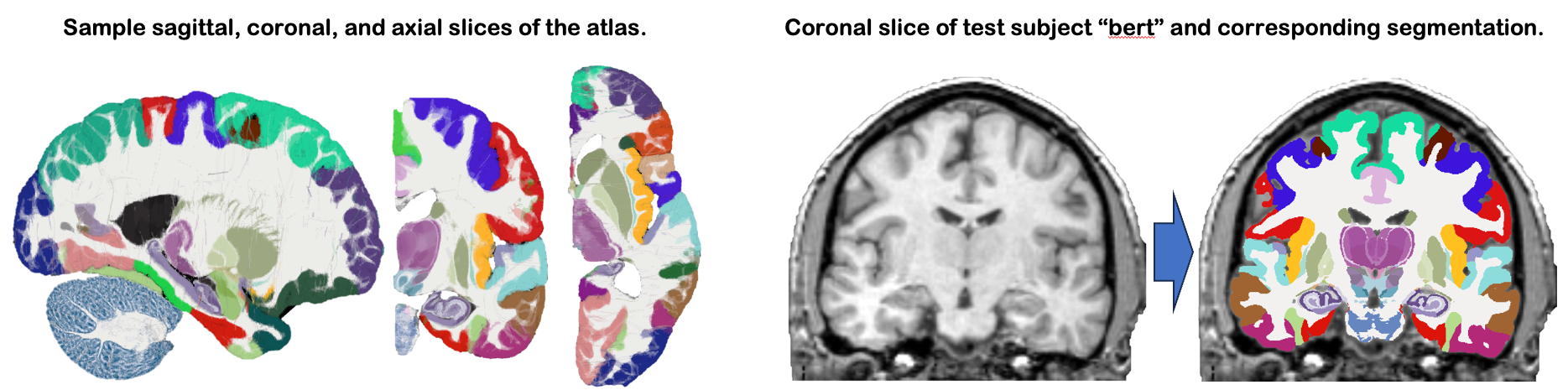
2. Installation
The first time you run this module, it will prompt you to download the atlas. Follow the instructions on the screen to obtain the atlas files.
In addition: this module calls mri_super_synth; if you have never used this command before, it will also prompt you to download a model file.
3. Basic usage
The entry point of the module is the command mri_histo_atlas_segment_fireants, which implements the fast version of the algorithm. This version relies on FireANTs (Jena et al) for fast nonlinear registration of the atlas. The command line is:
mri_histo_atlas_segment_fireants --i INPUT_SCAN --o OUTPUT_DIRECTORY --device [cpu/cuda] --side [left/right] --mode [invivo/cerebrum/hemi/exvivo]
INPUT_SCAN: scan to process, in mgz or nii(.gz) format.
OUTPUT_DIRECTORY: directory where segmentations, volume files, etc, will be created (more on this below).
DEVICE: set to cpu or cuda.
SIDE: left or right. If you're analyzing both sides, you're better off running them sequentially (rather than in parallel) since the SuperSynth preprocessing will be reused when processing the second hemisphere.
MODE: type of scan: invivo, exvivo, cerebrum (ex vivo without brainstem or cerebellum), hemi (ex vivo with single cerebral hemisphere).
4. Outputs
The output directory will contain the following files:
seg.[left/right].nii.gz: segmentation of left/right hemisphere
lut.txt: the lookup table to visualize seg.[left/right].nii.gz, for convenience
vols.[left/right].csv: files with volumes of the brain regions segmented by the atlas, in CSV format.
SuperSynth: directory with segmentation of the scan at the whole structure level.
Additional flags: if advanced options are used (more details below).
5. Advanced options
The code also accepts the following optional flags:
- --bf_mode: Decides the bias field basis function model. Options: dct (default), polynomial, hybrid.
- --write_rgb: Save an RGB image based on the posterior probabilites to disk.
- --write_bias_corrected: Save the bias corrected input image to disk.
- --device_registration: Define a different device for the registration. Can be used to save GPU memory when working with an GPU with limited memory. Options: cpu, cuda. Default is the same as --device.
- --threads: Control the number of cpu thread used to run the algorithm. Default value is -1, which uses all available threads.
- --skip: An integer skipping (downsampling) factor for estimating the model parameters. More skipping saves memory, but sacrifices accuracy. Default: 1 (no skipping).
- --resolution: The resolution of the output segmentation. By default 0.4mm, which is higher than the typical input scan, to reduce aliasing.
- --smoothing_steps_HRmask: Number of smoothing steps used when upsampling the 1mm brain mask from BrainFM. More smoothing makes the outer border less jagged, but too much smoothing reduces accuracy. Default: 3.
- --skip_bf: Skip the bias field correction. Can be used to save memory if the input scan is already bias corrected or does not have a bias field (non MRI modality).
--smooth_grad_sigma: Gradient field smoothing parameter for the nonlinear FireAnts registration. Default: 1.0.
--smooth_warp_sigma: Warp field smoothing parameter for the nonlinear FireAnts registration. Default: 0.25.
--optimizer_lr: Learning rate for the nonlinear FireAnts registration optimizer. Default: 0.5.
- --cc_kernel_size: Size of the window for calculating the cross-correlation registration metric. Default: 7.
- --rel_weight_labeldiff: Relative weight for the Dice loss metric in the nonlinear registration. Default: 2.5.
- --save_atlas_nonlinear_reg: Save the nonlinearly registered atlas. Default: false.
- --save_field: Save the nonlinear deformation field. Default: false.
- --save_jacobian: Save the Jacobian determinant (log10) of the deformation field. Default: false.
- --yaml_path: path of custom YAML files to define groups of ROIs
Some notes:
* If you are running out of memory, using --skip 2 can help without sacrificing much accuracy. * The defaults --smooth_grad_sigma 1 and --smooth_warp_sigma 0.25 are pretty liberal and can cope with massive deformation, e.g., as in the Hip-CT images shown in the paper "Fast segmentation with the NextBrain". If you are working with a population without very strong atrophy or deformation, you can multiply those values by 2 in order to get more regular atlas deformation fields (you can explore the deformation with the --save_jacobian option).
Also: you can flexibly change the groupings of the modeled structures using the .yaml files under the /data_simplified folder. The structure groupings for the Gaussian Mixture modeled are controlled by two files: gmm_components_fireants.yaml and combined_atlas_labels_fireants.yaml. Let's say, as an example, that you wanted to add the internal segment of globus pallidus (label 206) as its own structure. To model it separately, you would first create a new class, called e.g., Internal Segment Pallidum, in the combined_atlas_labels_fireants.yaml file, and list label 206 under that structure (while removing it from the pallidum class). Next, you would add the class, with exactly the same name, to the gmm_components_fireants.yaml file and decide how many Gaussian distributions should be used to model its intensities. To make the non-linear registration aware of the contrast, you would add the structure, again with exactly the same name, to the file called recipe_intensities_cheating_image_fireants.yaml, and decide how its intensity should be generated from the seven structures than can be always reliably segmentation using BrainFM (see the file for examples).
6. Frequently asked questions (FAQ)
I have an ex vivo hemisphere with cerebellum and/or brainstem
If you use the hemi mode, you will not get the cerebellum or brainstem. Use the exvivo mode instead (with the caveat that you may lose some voxels around the medial wall, which may get assigned to the contralateral hemisphere).
Can the exvivo model handle arbitrary orientations of the input
No, it cannot. You need to manually reorient the brain to RAS (e.g., with Freeview).
Do I need a GPU?
Certainly not! The code should run in less than half an hour on any semi-modern workstation, if you allocate enough threads (or about two hours for an ex vivo scan at 0.25mm resolution).
What happened to the "full Bayesian" and "SynthMorph" versions?
To simplify the codebase, we are focusing on this method, which is fast but also versatile in terms of modeling / registration (as opposed to SynthMorph).
