|
Size: 12915
Comment:
|
Size: 13163
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 166: | Line 166: |
=== Q: How can I get measurements of the lobes (parietal, temporal, frontal, & occipital)? === A: You can either use the PALS_B12 atlas (mapped to fsaverage in v5.0) or create your own following the directions on the CorticalParcellation wiki. |
|
| Line 167: | Line 171: |
FAQ
- The purpose of this FAQ is to provide a Wiki space where users can add the most frequent questions to avoid asking already answered questions in the support list.
Contents
- General
-
Computing
- Q. How long does it take to finish a reconstruction?
- Q. How can I reduce the time of recon-all in a group of patients?
- Q. I want to use freesurfer to quantity cortical thickness, and I want to use the software under Linux. What equipment for computer do you recommend?
- Q. Is it possible to run FreeSurfer in Ubuntu Linux?
-
Processing/Re-processing Data in FreeSurfer
- Q. How to run recon-all in a ssh terminal
- Q. I had a [Power Outage|Computer Failure|Spilled coffee on computer|etc], how can I resume the recon-all?
- Q. I have already skull-stripped data. Can I submit it to recon-all?
- Q. I made white matter and pial edits to my volume. Do I need to run -autorecon2-wm and when that is finished run -autorecon2-pial?
- Common Error messages
-
FreeSurfer Output Questions
- Q. The surfaces near the medial wall, hippocampus, and amygdala aren't accurately following the structures there. How can I fix this?
- Q: It seems that there is a 5mm thickness upper limit in my volumes, is it normal? How can I change this?
- Q. Why in many subjects the insular cortical surface seems so thick? Is the convoluted nature of the Insula that causes that?
- Q. What are the sulc and curv overlays (in QDEC) showing us?
- Q. How would I put a FreeSurfer output volume and/or segmentation back into the same space as my original anatomical input (native space)?
-
Analysis of FreeSurfer Data
- Q. I am trying to measure the cortical thickness of specific ROI how can I do this?
- Q. I am using Qdec to examine the anatomical differences between two groups of subjects. The surface-based measures I can select are thickness, area, area.pial, sulc, curv, and jacobian_white. Could anybody tell me what anatomical features the later three (sulc, curv, and jacobian_white) actually measure?
- Q: How can I get measurements of the lobes (parietal, temporal, frontal, & occipital)?
- FreeSurfer & Matlab
General
Q. How can I help this FAQ?
A: If you are able to edit pages go ahead! If you don't have permission to write on the Wiki send an e-mail to ppj at netfilter dot com dot br
Q. What are the advantages of FreeSurfer over VBM?
A:
- FS uses geometry to do inter-subject registration, which experience has shown results in a much better matching of homologous cortical regions than volumetric techniques.
- FS allows you to look at the two components of volume separately (thickness and surface area). It has been found that these two do not necessarily track one another, and in the worst case where one is increasing and the other decreasing the volume change can be 0.
- The target that FS uses for registration (the white matter surface geometry) is completely invariant to gm atrophy, so gm changes won't result in a different registration.
Q. How do I know the version of FreeSurfer I'm running?
A: Run:
- recon-all -version
Computing
Q. How long does it take to finish a reconstruction?
A: It depends on your processor speed and machine performance (notice that dual/quad/hex core or hyperthead won't speed up one analysis significantly), give a look at the link below in the section "Step-wise directives": http://surfer.nmr.mgh.harvard.edu/fswiki/recon-all
Also you can contribute to our running time statistics: https://surfer.nmr.mgh.harvard.edu/fswiki/ReconAllRunTimes
Q. How can I reduce the time of recon-all in a group of patients?
A: FreeSurfer run its process in a non-parallel environment, so you won't have benefit from a dual/quad/hex core machine for a single case analysis. However if you have many cases you can start two FreeSurfer recon-all process in the same machine and theoretically you can reduce by half the time to analyze your group of cases. A similar procedure can also be used in quad/hex-core environment. Note that to take benefit of a multi-core environment you need to use a SMP kernel in your OS.
Q. I want to use freesurfer to quantity cortical thickness, and I want to use the software under Linux. What equipment for computer do you recommend?
This answer requires constant updates:
For Best Performances:
Use a multi-Core processor Intel (a i7-900 series processor or Xeon 6500/7500 Series)
Install at Least 8GB of Memory
If you prefer AMD you can use:
AMD Opteron Six-Core or Phenon II X6
Other alternative much more expensive is: Mac Pro with 12 cores and 16GB RAM.
The number of cores is roughly the number of studies you can process simultaneously. Notice that each process will take 20-24 hrs.
Keep in mind that you need to use the fastest memory in order to achieve maximum benefit from multi-core architecture.
Q. Is it possible to run FreeSurfer in Ubuntu Linux?
A: Yes. Ubuntu Linux is basically a Debian distro, so you should use FreeSurfer RH9 version. Depending on your video card you should disable the DRI using the option NoDRI in the Device section of your X configuration file. Notice that in older Ubuntu version there's a bug that prevents NoDRI from working.
Processing/Re-processing Data in FreeSurfer
Q. How to run recon-all in a ssh terminal
A: This is tricky, because recon-all spawns many process and redirect I/O to the terminal that opened it. Run
recon-all -s subjid -all &> /dev/null &
disown -a
Q. I had a [Power Outage|Computer Failure|Spilled coffee on computer|etc], how can I resume the recon-all?
A: You can try
recon-all -make all -s subject
Also, depending on your version, there might be a file in the scripts directory called "IsRunning". You should delete this if it is there.
Q. I have already skull-stripped data. Can I submit it to recon-all?
A: If your skull-stripped volume does not have the cerebellum, then no. If it does, then yes, however you will have to run the data a bit differently.
First you must run only -autorecon1 like this:
recon-all -autorecon1 -noskullstrip -s <subjid>
Then you will have to make a symbolic link or copy T1.mgz to brainmask.auto.mgz and a link from brainmask.auto.mgz to brainmask.mgz. Finally, open this brainmask.mgz file and check that it looks okay (there is no skull, cerebellum is intact; use the sample subject bert that comes with your FreeSurfer installation to make sure it looks comparable). From there you can run the final stages of recon-all:
recon-all -autrecon1 -autorecon3 -s <subjid>
Q. I made white matter and pial edits to my volume. Do I need to run -autorecon2-wm and when that is finished run -autorecon2-pial?
A: No. If you made both white matter and pial edits, you only need to run -autorecon2-wm which is higher up in the processing stream (before the steps of -autorecon2-pial). By running -autorecon2-wm, it will perform the steps necessary to fix a white matter edit and then all the remaining steps in -autorecon2 including those steps that -autorecon2-pial would run. The OtherUsefulFlags wiki shows the hierarchy of the -autorecon2 flag shortcuts (-autorecon2-cp starts before -autorecon2-wm which starts before -autorecon2-pial). In general, you should start the processing stream at the earliest step where you made an manual intervention and the rest will be taken care of. Don't forget to run -autorecon3!
Common Error messages
Q. Help! I got this error message: "mri_watershed error: GLOBAL region of the brain empty!" - what should I do?
A: Run this:
recon-all -skullstrip -no-wsgcaatlas -s <subjid>
If that goes through without error, you can finish processing your data by running:
recon-all -autorecon2 -autorecon3 -s <subjid>
Q. I get an error message from the Talairach Failure Detection. What does this mean?
A: In certain versions of FreeSurfer, the talairach failure detection is too conservative so this may not be a problem. First, you will want to check the talairach transform to make sure it looks okay. Directions on how to do that are here: http://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/Talairach
Essentially, you want to use the command:
tkregister2 --mgz --s <subjid> --fstal
and see if the green lines line up with the anatomy of the blurry brain image. If so, then you can run this subject as you did before, just add -notal-check to your command. For example, recon-all -all -s subjid -notal-check. If the green lines do not line up well with the anatomy, you will want to follow directions on the wiki page mentioned above to fix it.
FreeSurfer Output Questions
Q. The surfaces near the medial wall, hippocampus, and amygdala aren't accurately following the structures there. How can I fix this?
A: The good news is you don't have to! These areas are generally unreliable for a thickness study so you would want to exclude them from your analysis. If you use glmfit, it calls the ?h.cortex.label file in order to determine thickness. This file automatically has set these areas to zero so they will not be included in your analysis. To see what area ?h.cortex.label excludes you can load it in tksurfer on top of the inflated surface. If you have tkmedit open at the same time for the same subject, you can use the Go To Saved Point function to make sure areas of concern are in this excluded section. *Note: If you are using a FreeSurfer version older than 4.3.0, the excluded region includes the insula. If you are using an analysis program other than glmfit, just be sure to use the ?h.cortex.label file to get your thickness measurements.
The picture below shows the ?h.cortex.label (outlined in yellow) over the ?h.aparc.annot. Notice how the ?h.cortex.label includes some of the ?h.unknown.label:
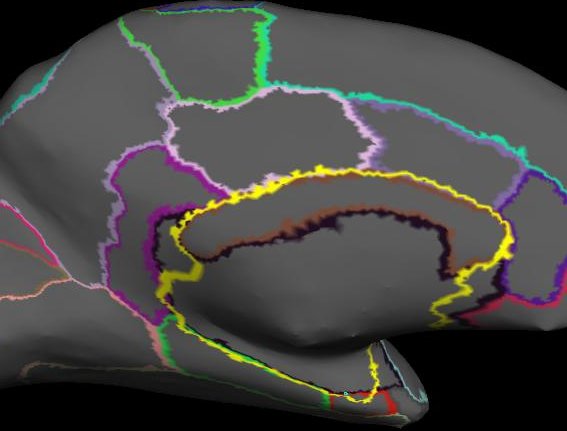
As always, use the example subject bert that comes with the FreeSurfer installation to see what we define as acceptable surfaces.
Q: It seems that there is a 5mm thickness upper limit in my volumes, is it normal? How can I change this?
A: We did this to prevent noncortical regions such as the basal ganglia from corrupting the thickness measure through averaging. With the ?h.cortex.label it is probably no longer needed, however you can use mris_thickness -max <max thick> to generate a thickness with a different max. Take a look in the example below:
mris_thickness -max 10 bert lh newlh.thickness
The file newlh.thickness will be created inside the surf directory of your subject. But we don't think the true thickness is ever that much except in pathological cases like dysplasia and other disorders of cortical development.
Q. Why in many subjects the insular cortical surface seems so thick? Is the convoluted nature of the Insula that causes that?
A: It's not the convoluted nature of the Insula. It's the fact that extreme capsule is so thin that it frequently isn't very apparent on MR, and so there appears to be continuous gray matter from the basal ganglia into the cortex. We think version 4.0 fixes this.
Q. What are the sulc and curv overlays (in QDEC) showing us?
A: The 'sulc' conveys information on how far removed a particular vertex point on a surface is from a hypothetical "mid-surface" that exists between the gyri and sulci. This surface is chosen so that the "mean" of all these displacements is zero. The 'sulc' gives a indication then of linear distance and displacements: how "deep" and how "high" are brain folds
In FreeSurfer, gyri have negative 'sulc' values, are colored green, and indicate how far "down" a point has to travel to reach this "mid-surface". Sulci have positive 'sulc' values, are colored red, and indicate how far "up" a point needs to travel to reach the mid-surface.
The 'curv' conveys information on the curvature (not distance) at a specific vertex point. The color conveys the sign, and is just an arbitrary choice. The sharper the curve, the higher the value (positive or negative). Areas with positive curvature, are colored red, and correspond to curvatures in sulci, i.e. curving "up". Areas with negative curvature are colored green, and correspond to curves pointing "down", i.e. gyri.
So, in a nutshell, the difference is that the 'curv' files contain information about curvatures, and the 'sulc' files contain information about displacement.
Q. How would I put a FreeSurfer output volume and/or segmentation back into the same space as my original anatomical input (native space)?
A: Please see directions on how to do this here.
Analysis of FreeSurfer Data
Q. I am trying to measure the cortical thickness of specific ROI how can I do this?
A: You can save the ROI to a label file and then use:
mris_anatomical_stats -l <label file>
Q. I am using Qdec to examine the anatomical differences between two groups of subjects. The surface-based measures I can select are thickness, area, area.pial, sulc, curv, and jacobian_white. Could anybody tell me what anatomical features the later three (sulc, curv, and jacobian_white) actually measure?
sulc = "average convexity" from our 1999 reconII paper(https://surfer.nmr.mgh.harvard.edu/ftp/articles/fischl99b-recon2.pdf).
Essentially measures the depth/height of each point above the average surface.
curv = smoothed mean curvature.
jacobian_white = the jacobian of the spherical transform. Measures the amount of distortion needed to warp a subject into register with the atlas.
You might want to look at the slides downloadable from the top of this page:
http://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial
which contain pictures showing the meaning of 'curv' and 'sulc'.
Q: How can I get measurements of the lobes (parietal, temporal, frontal, & occipital)?
A: You can either use the PALS_B12 atlas (mapped to fsaverage in v5.0) or create your own following the directions on the CorticalParcellation wiki.
FreeSurfer & Matlab
Q. How can I make an histogram of cortical thickness?
A: In Matlab you can use the read_curv() function, as in the example below.
thick = read_curv ('/usr/local/freesurfer/subjects/bert/surfer/lh.thickness');
hist (thick,100);
Notice that there are many zero values that refer to areas where there's no cortical surface. The best procedure is to create a new array without the zero values and then make the histogram.
