- If preferred, one can use the edited cross sectional data to create a new base. However, edits made to the brain.finalsurfs volume of the cross sectional data sets will not transfer to the base since the base creation takes off from the norm.mgz of the cross data sets.
Therefore, in order to fix the base, one must edit the brain.finalsurfs volume of the base as follows:
A. Below is a visualization of the problem area in the base: brain.finalsurfs.mgz, slice 89 at coordinates 98, 172, 89 in right hemi. The problem area extends through slices 85 - 94.
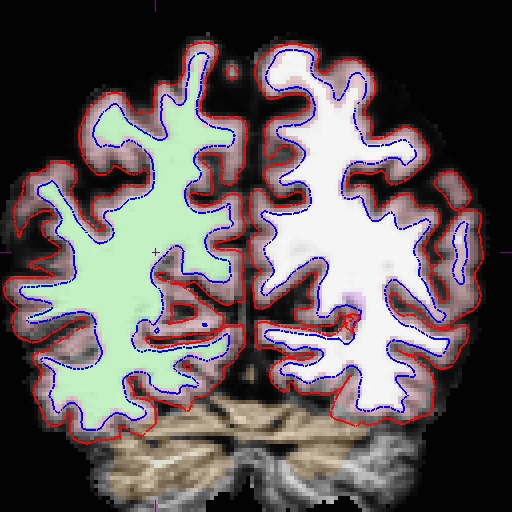
And here is the area zoomed in:
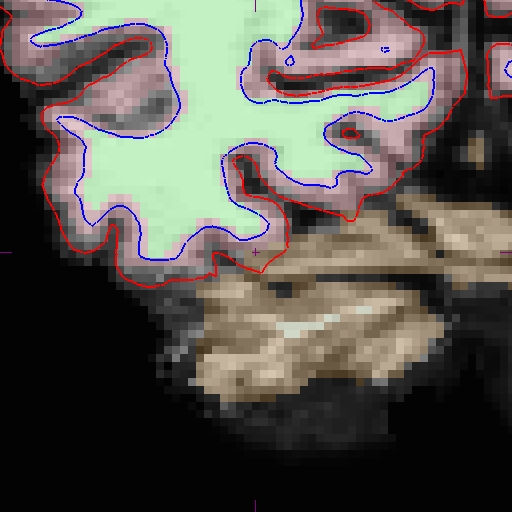
Next, edit the affected slices by removing the appropriate voxels, and save the changes to the aux volume. Then, cd into the mri directory of the base data set, and make a copy of the edited brain.finalsurfs.mgz file to one called brain.finalsurfs.manedit.mgz.: 1. cd OAS2_0002/mri 2. cp brain.finalsurfs.mgz brain.finalsurfs.manedit.mgz
Next, run the following to have edits take effect: {{{recon-all -autorecon3-pial -base OAS2_0002 }}}
Results will look as follows:
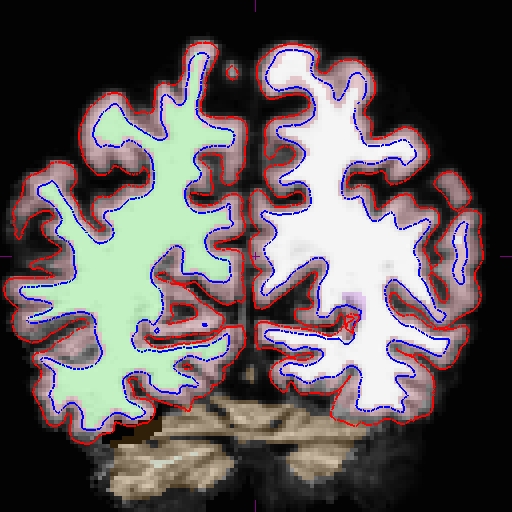
Here is zoomed in version of above:
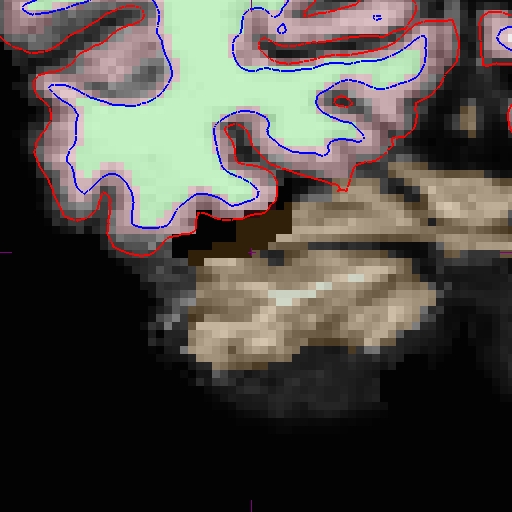
Now that the base and cross sectionals are fixed, you must re-create the longitudinal data sets for each time point. Return to main page to read that section.
