SynthSR
This functionality was made available in version 7.3.0 of FreeSurfer.
Please note that version 2.0 of the tool was released in the development version on February 8th 2023.
Please note that we have a new tool (SuperSynth) that: does not inpaint lesions; estimates T1, T2, and FLAIR contrast; and is comptatible with ex vivo scans and single hemispheres.
Author: Juan Eugenio Iglesias
E-mail: jiglesiasgonzalez [at] mgh.harvard.edu
Rather than directly contacting the author, please post your questions on this module to the FreeSurfer mailing list at freesurfer [at] nmr.mgh.harvard.edu
If you use this tool in your analysis, please cite:
Joint super-resolution and synthesis of 1 mm isotropic MP-RAGE volumes from clinical MRI exams with scans of different orientation, resolution and contrast. JE Iglesias, B Billot, Y Balbastre, A Tabari, J Conklin, RG Gonzalez, DC Alexander, P Golland, BL Edlow, B Fischl, for the ADNI. Neuroimage, 118206 (2021).
SynthSR: a public AI tool to turn heterogeneous clinical brain scans into high-resolution T1-weighted images for 3D morphometry. JE Iglesias, B Billot, Y Balbastre, C Magdamo, S Arnold, S Das, B Edlow, D Alexander, P Golland, B Fischl. Science Advances, 9(5), eadd3607 (2023).
If you use the Hyperfine version, please cite this paper as well:
Quantitative Brain Morphometry of Portable Low-Field-Strength MRI Using Super-Resolution Machine Learning. JE Iglesias, R Schleicher, S Laguna, B Billot, P Schaefer, B McKaig, JN Goldstein, KN Sheth, MS Rosen, WT Kimberly. Radiology, 220522 (2022).
Contents
- Motivation and General Description
- Installation
- Usage
- Processing low-field scans (e.g., Hyperfine)
- Processing CT scans
- Old multispectral (T1+T2) model for Hyperfine scans
- Frequently asked questions (FAQ)
1. Motivation and General Description
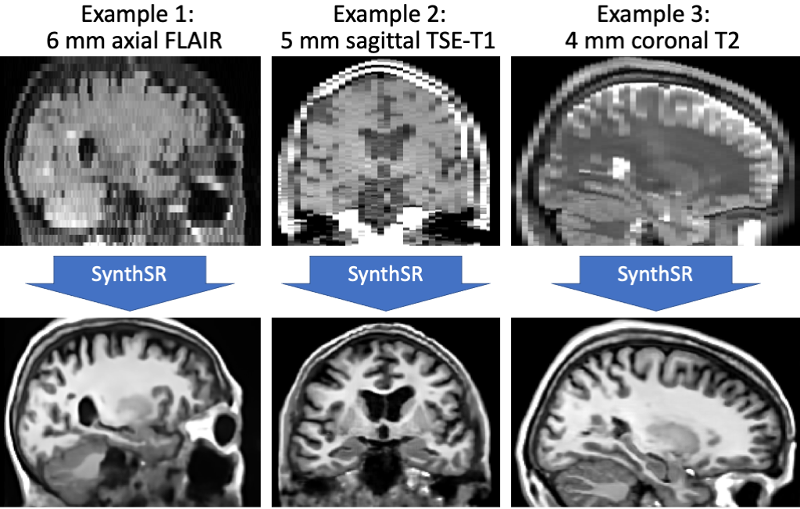
This tool implements SynthSR, a convolutional neural network that turns a clinical MRI scan (or even CT scan!) of any orientation, resolution and contrast into 1 mm isotropic MP-RAGE, while inpainting lesions (which enables easier segmentation, registration, etc). You can then run your favorite neuroimaging software (including FreeSurfer, of course!) on these synthetic images for segmentation / registration / any other analysis.
2. Installation
The first time you run this module, it will prompt you to install Tensorflow. Simply follow the instructions in the screen to install the CPU or GPU version.
If you have a compatible GPU, you can install the GPU version for faster processing, but this requires installing libraries (GPU driver, Cuda, CuDNN). These libraries are generally required for a GPU, and are not specific for this tool. In fact you may have already installed them. In this case you can directly use this tool without taking any further actions, as the code will automatically run on your GPU.
3. Usage
We provide an "all purpose" model that can be applied to a scan of any resolution of contrast. Once FreeSurfer has been sourced, you can simply test SynthSR on your own data with:
mri_synthsr --i <input> --o <output> --threads <n_threads> [--v1] [--lowfield] [--ct]
where:
<input>: path to an image to super-resolve / synthesize. This can also be a folder, in which case all the image inside that folder will be processed.
<output>: path where the synthetic 1 mm MP-RAGE will be saved. This must be a folder if --i designates is a folder.
<n_threads>: (optional) number of CPU threads to use. The default is just 1, so crank it up for faster processing if you have multiple cores!
--v1: (optional) use version 1.0 of SynthSR (model from July 2021); this is helpful for reproducibility purposes.
--lowfield: (optional) use this flag when processing low-field scans with limited resolution and signal-to-noise ratios (see below).
--ct: (optional) use this flag when processing CT scans (details below).
The synthetic 1mm MP-RAGE will be of a standard contrast, bias field corrected, and with white matter lesions inpainted.
4. Processing low-field scans (e.g., Hyperfine)
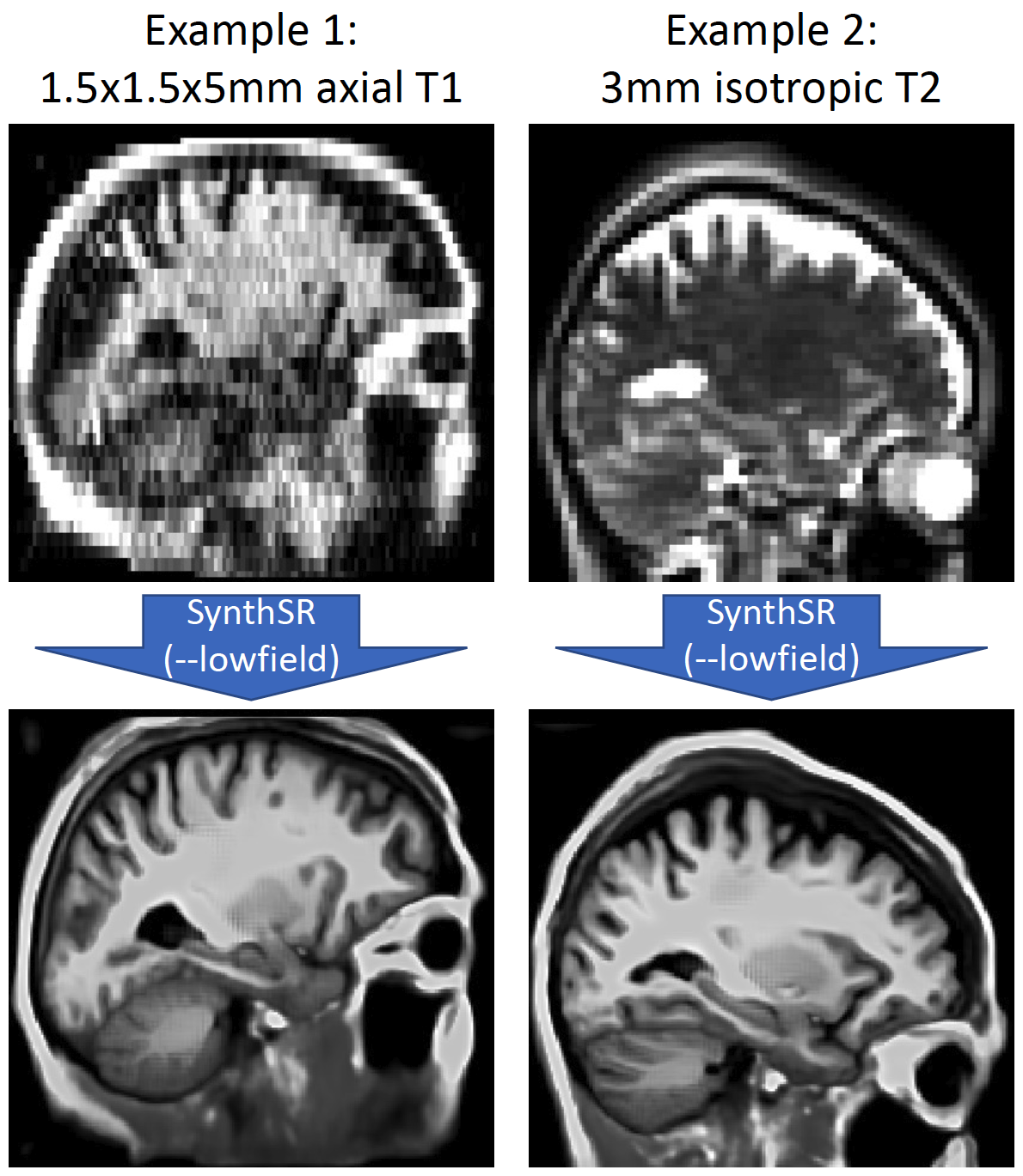
In the 1.0 version of SynthSR, we provided a dedicated, multispectral (T1+T2) model for Hyperfine scans. This was problematic for two reasons: the pulse sequence requirements, and the alignment between the two modalities. While this model is still provided for reproducibility purposes (see below), we now have a single-scan model for low-field data. This is very similar to the "general" model, but the data synthesized during training has less resolution, more noise, and starker signal dropout. Below are two examples, from a 1.5x1.5x5mm axial T1 and from a 3mm isotropic T2:
5. Processing CT scans
Regarding CT scans: SynthSR does a decent job with CT ! The only caveat is that the dynamic range of CT is very different to that of MRI, so they need to be clipped to [0, 80] Hounsfield units. You can use the --ct flag to do this, as long as your image volume is in Hounsfield units. If not, you will have to clip to the Hounsfield equivalent yourself (and not use --ct).
6. Old multispectral (T1+T2) model for Hyperfine scans
In version 1.0, we provided a dedicated, multispectral (T1+T2) model for low-field scans acquired with the "standard" Hyperfine sequences (1.5x1.5x5mm axial). This model is still available, for reproducibility purposes. You can used it with the command:
mri_synthsr_hyperfine --t1 <t1> --t2 <t2> --o <output> --threads <n_threads>
where, as in the previous version, <t1>, <t2> and <output> can be single files or directories.
If there is motion between the T1 and T2 scans, the T2 needs to be pre-registered to the space of the T1, but without resampling to the 1.5x1.5x5mm space of the T1, which would introduce large resampling artifacts. This can be done with FreeSurfer's mri_robust_register:
mri_robust_register --mov T2.nii.gz --dst T1.nii.gz --mapmovhdr T2.reg.nii.gz --cost NMI --noinit --nomulti --lta /dev/null
7. Frequently asked questions (FAQ)
Does running this tool require preprocessing of the input scans?
No! Because we applied aggressive augmentation during training (see paper), this tool is able to segment both processed and unprocessed data. So there is no need to apply bias field correction, skull stripping, or intensity normalization.
What formats are supported ?
This tool can be run on Nifti (.nii/.nii.gz) and FreeSurfer (.mgz) scans.
How can I increase the speed of execution?
If you have a multi-core machine, you can increase the number of threads with the --threads flag (up to the number of cores).
How can I take advantage of multiple acquisitions when available (e.g., T1 + T2 + FLAIR)?
We have found that the positive effect of having multiple MRI modalities available is offset by the registration errors between the channels. If you have multiple usable acquisitions, we recommend that you run SynthSR + downstream analyses in all of them and average (or take median) of the results. For example: if you are interested in computing the hippocampal volume with FSL, you could run SynthSR on all the scans, then the FSL segmentation on all the synthetic scans, and average the results.
Why aren't the predictions perfectly aligned with their corresponding images?
This is probably of problem of image viewer! Indeed, this issue might arise when performing super-resolution, when the input images and their predictions are not at the same resolution, and some viewers cannot cope with resolution changes. We recommend using FreeView (shipped with FreeSurfer), which is resolution-aware and does not have these problems.
