|
Size: 3427
Comment:
|
Size: 3876
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 13: | Line 13: |
| * [[https://www.sciencedirect.com/science/article/pii/S1053811921004833|Joint super-resolution and synthesis of 1 mm isotropic MP-RAGE volumes from clinical MRI exams with scans of different orientation, resolution and contrast]]. JE Iglesias, B Billot, Y Balbastre, A Tabari, J Conklin, RG Gonzalez, DC Alexander, P Golland, BL Edlow, B Fischl, for the ADNI. Neuroimage, 118206 (2021). | * [[https://www.sciencedirect.com/science/article/pii/S1361841523000506|SynthSeg: Segmentation of brain MRI scans of any contrast and resolution without retraining]]. B Billot, DN Greve, O Puonti, A Thielscher, K Van Leemput, B Fischl, AV Dalca, JE Iglesias. Medical Image Analysis, 83, 102789 (2023). * [[https://www.pnas.org/doi/10.1073/pnas.2216399120|Robust machine learning segmentation for large-scale analysis of heterogeneous clinical brain MRI datasets]]. B Billot, C Magdamo, SE Arnold, S Das, JE Iglesias. PNAS, 120(9), e2216399120 (2023). |
| Line 21: | Line 22: |
| * SynthSeg: to obtain an aseg.auto_noCCseg.mgz and to compute a Talairach transform | * SynthSeg: to obtain a volumetric segmentation and linear registration to Talairach space |
| Line 23: | Line 24: |
| * SynthSurfaces: to fit surfaces by predicting the distance maps and reconstructing topologically accurate cortical surfaces | * SynthDist: to fit surfaces by predicting the distance maps and reconstructing topologically accurate cortical surfaces |
| Line 43: | Line 44: |
| ''This stream runs a bit faster than the original recon-all, since the volumetric segmentation is much faster than the iterative Bayesian method in the standard stream'' | ''This stream runs a bit faster than the original {{{recon-all}}}, since the volumetric segmentation is much faster than the iterative Bayesian method in the standard stream'' === Outputs: === ''This stream will create a directory structure that is almost the same as {{{recon-all}}}, but with some minor differences in the {{{SUBJECT_DIR/mri}}}'' - Rest of the directories are the same with the different parcellations supported by FreeSurfer. |
recon-all-clinical
This functionality is now available in the developer version of FreeSurfer.
Author: Karthik Gopinath
E-mail: kgopinath[at]mgh[dot]harvard[dot]edu
Please post your questions on this module to the FreeSurfer mailing list at freesurfer[at]nmr.mgh.harvard.edu rather than directly contacting the author.
If you use this package in your analysis, please cite:
Cortical analysis of heterogeneous clinical brain MRI scans for large-scale neuroimaging studies. K Gopinath, DN Greeve, S Das, S Arnold, C Magdamo, JE Iglesias
SynthSeg: Segmentation of brain MRI scans of any contrast and resolution without retraining. B Billot, DN Greve, O Puonti, A Thielscher, K Van Leemput, B Fischl, AV Dalca, JE Iglesias. Medical Image Analysis, 83, 102789 (2023).
Robust machine learning segmentation for large-scale analysis of heterogeneous clinical brain MRI datasets. B Billot, C Magdamo, SE Arnold, S Das, JE Iglesias. PNAS, 120(9), e2216399120 (2023).
SynthSR: a public AI tool to turn heterogeneous clinical brain scans into high-resolution T1-weighted images for 3D morphometry. JE Iglesias, B Billot, Y Balbastre, C Magdamo, S Arnold, S Das, B Edlow, D Alexander, P Golland, B Fischl. Science Advances, 9(5), eadd3607 (2023).
General description:
This tool performs recon-all-clinical, the first out-of-the-box cortical surface reconstruction and analysis of brain MRI scans of any modality, contrast and resolution without retraining and fine-tuning.
This "Recon-all-like" stream for clinical scans of arbitrary orientation/resolution/contrast is essentially a combination of:
SynthSeg: to obtain a volumetric segmentation and linear registration to Talairach space
SynthSR: to have a higher resolution 1mm MPRAGE for visualization
SynthDist: to fit surfaces by predicting the distance maps and reconstructing topologically accurate cortical surfaces
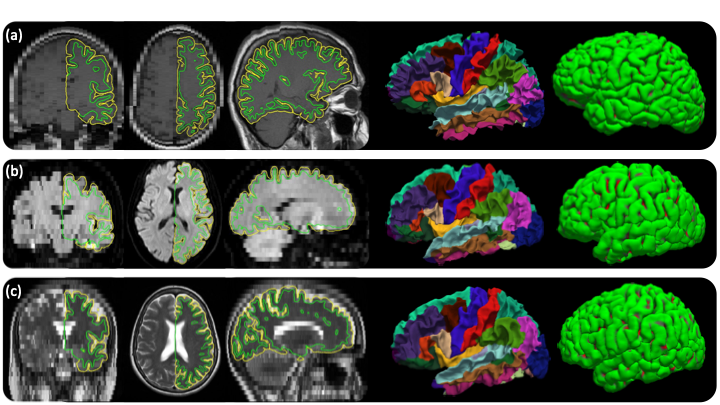
Out of the box cortical surface reconstruction and analysis of heterogenous scans. (a)Sagittal T1 scan with .4×.4×6mm resolution. (b)Axial FLAIR scan with 1.7×1.7×6mm resolution. (c)Axial T2-weighted scan with .9×.9×6mm resolution. The WM surface with cortical parcellation overlaid and pial surfaces are also shown.
Usage:
OnceFreeSurfer has been sourced, you can simply run recon-all-clinical on your own data with
recon-all-clinical.sh INPUT_SCAN SUBJECT_ID THREADS [SUBJECT_DIR]
where:
- INPUT_SCAN: path to an image that will be processed.
- SUBJECT_ID: specifies the name or ID of the subject you would like to use. A directory with that name will be created for all the subject's FreeSurfer output.
- THREADS (optional): number of CPU threads to use. The default is just 1, so crank it up for faster processing if you have multiple cores!
- SUBJECT_DIR: only necessary if the environment variable SUBJECTS_DIR has not been set when sourcing FreeSurfer or if you want to override it.
This stream runs a bit faster than the original recon-all, since the volumetric segmentation is much faster than the iterative Bayesian method in the standard stream
Outputs:
This stream will create a directory structure that is almost the same as recon-all, but with some minor differences in the SUBJECT_DIR/mri - Rest of the directories are the same with the different parcellations supported by FreeSurfer.
