Correcting topological defects
To follow this exercise exactly, make sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names.
The automatic topology fixer is used to make sure that there are no "holes" or "handles" in the surface (ie, that it has the topology of a closed sphere). These topology errors are actually due to white matter segmentation errors, ie, true white matter being labeled as non-white matter or non-white matter being labeled as white matter. Sometimes, the automatic topology fixer run during recon-all fails. Failures can lead the white matter (WM) surface (yellow line) to exclude some voxels that are white matter or to include some voxels that are not white matter. Some times, the topological defect can be very large (eg, cerebellum not removed); large defects can cause recon-all to appear to hang or crash after printing something like CORRECTING DEFECT...). Simple edits to wm.mgz volume can correct these problems.
Editing the Volume
Subject topo_defect_before is an example of topological defect and this page will walk you through fixing it. First, make sure you have topo_defect_before open in freeview (you may have already run this command).
freeview -v topo_defect_before/mri/brainmask.mgz \ topo_defect_before/mri/wm.mgz:colormap=heat:opacity=0.4:visible=0 \ -f topo_defect_before/surf/lh.white:edgecolor=yellow \ topo_defect_before/surf/lh.pial:edgecolor=red \ topo_defect_before/surf/rh.white:edgecolor=yellow \ topo_defect_before/surf/rh.pial:edgecolor=red \ topo_defect_before/surf/lh.orig.nofix:visible=0
Note: you only need the ?h.orig.nofix surface here; the other surfaces might not have been created if recon-all failed due to large topology defects. Use the PageUp and PageDown keys to scroll through the volume (or the up and down arrows) to find the error in white matter surface. If you look at coronal slice 57, there is a defect in the left hemisphere closer to the midline, where some white matter voxels are segmented incorrectly. This type of topological defect is called a 'hole' which needs to be filled as opposed to a handle which needs to be cut.
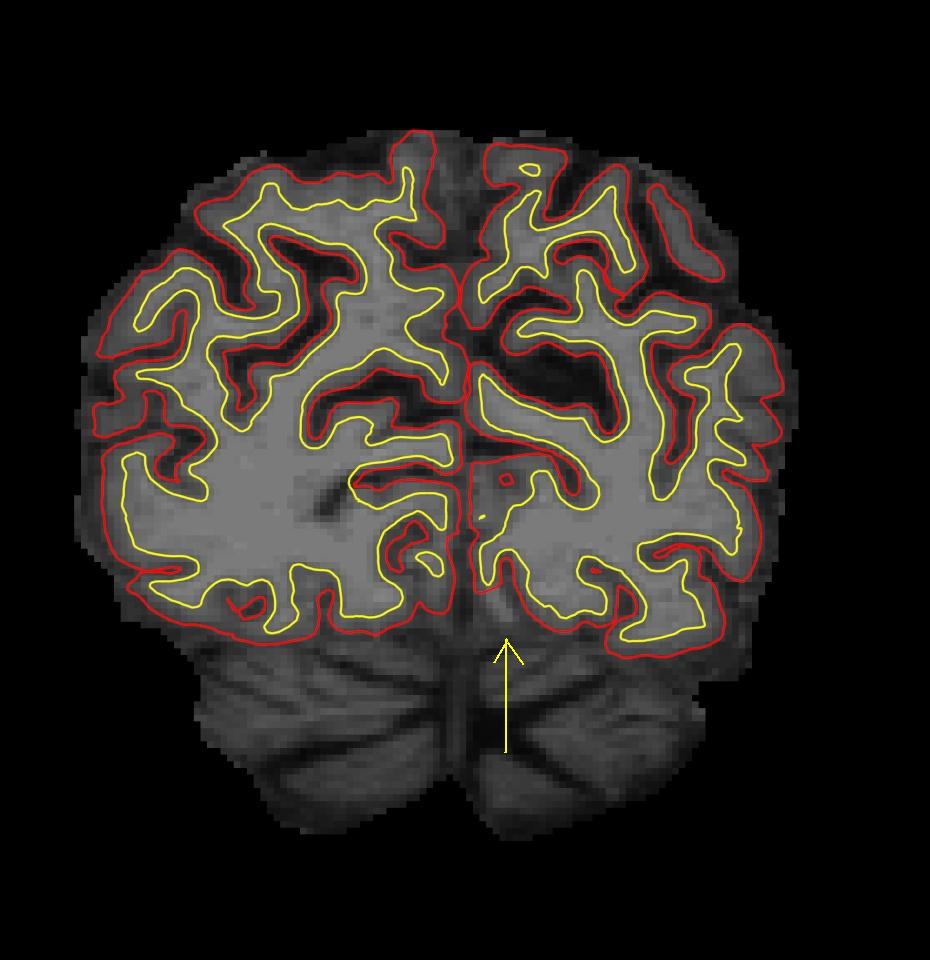
Now make the wm.mgz volume visible by clicking on the checkbox next to the wm volume in Freesurfer. As you can see, some white matter voxels are excluded from the white matter surface. This is because there is a "break" in the white matter mask (red filled in volume). Some voxels that are clearly part of the white matter are not colored red, causing the white matter surface to not include all voxels in this area. If you are having trouble finding the break in the wm surface, try looking at the lh.orig.nofix surface in the coronal plane instead. This surface will follow voxel boundaries and will highlight more clearly which white matter voxels have been excluded.
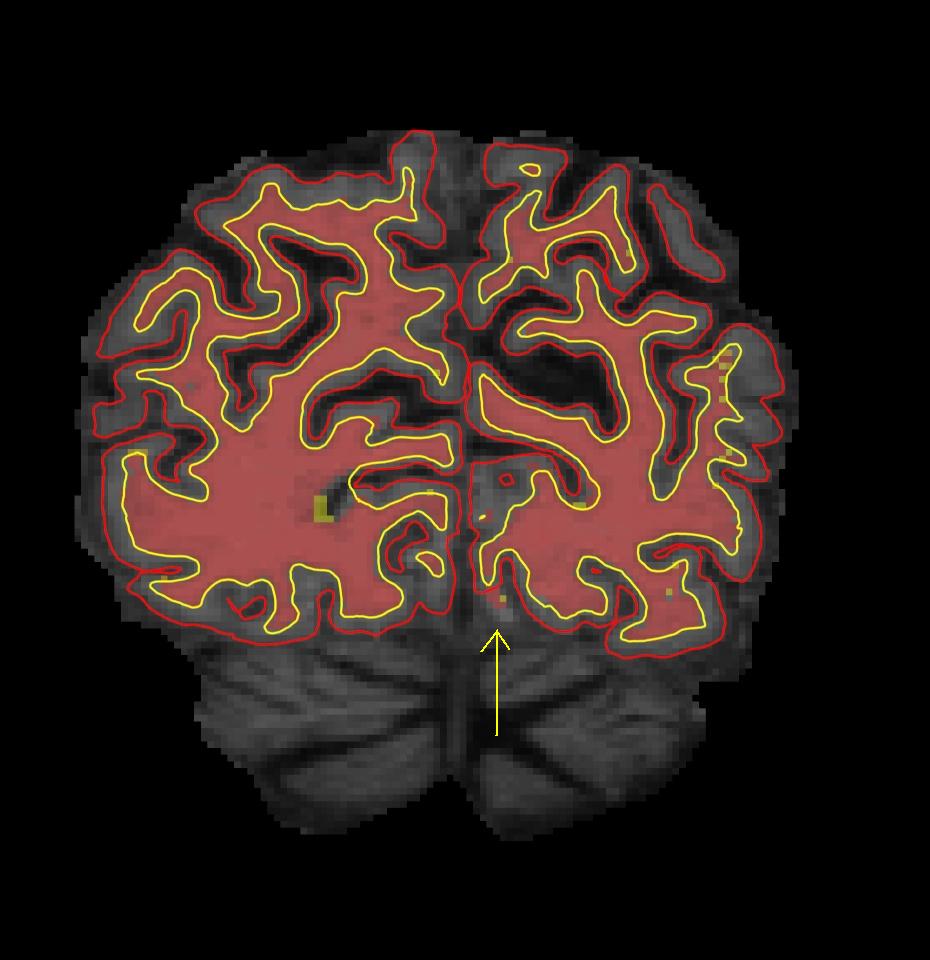
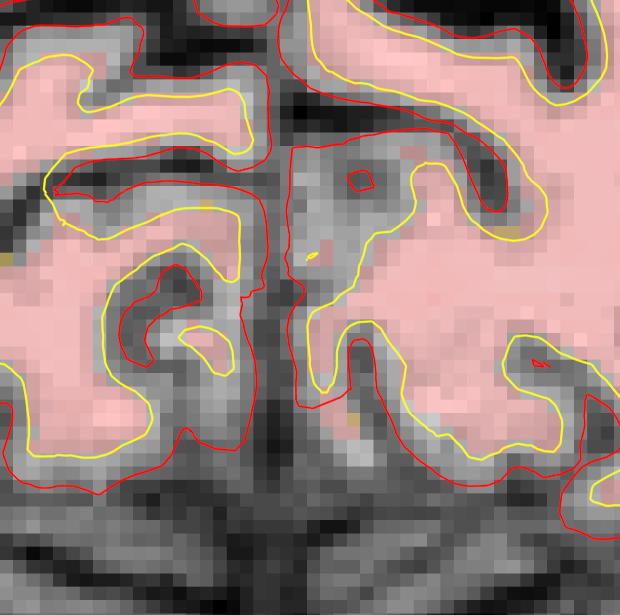
To view the defect as part of the surface, switch to the 3D view and check the box corresponding to the lh.smoothwm.nofix surface, and uncheck all the other surfaces to prevent any overlap. This surface is the same as the orig.nofix surface, just smoothed so that it will not follow voxel boundaries. If you're interested in seeing the difference, you can also view the orig.nofix surface in Freeview. However, you will not see the defect in the wm surface in 3D view. This is because orig.nofix and smoothwm.nofix are thought of as "rough drafts" of the final white surface. After smoothwm.nofix is created, the surface still undergoes further processing to become the final white surface. This further processing includes the automatic topology fixer, which did not completely correct this topological defect. If your view of the surfaces is obstructed by the 2D slices appearing in the 3D view, you can shut them off by pressing Ctrl-Shft-S on the keyboard or by clicking the View option in Freeview and then clicking on Show Slices (3D View).
Below are a couple of zoomed-in images of lh.smoothwm.nofix rendering the surface as a mesh, exposing all the vertices which create the surface topology. By default, surfaces opened in freeview are not rendered as meshes. To enable the mesh rendering, click a surface in the sidebar, and select Surface & mesh under the Render drop-down menu. You can see the error as pointed by the yellow arrow, which is a hole that needs to be filled to correct this type of topological defect.
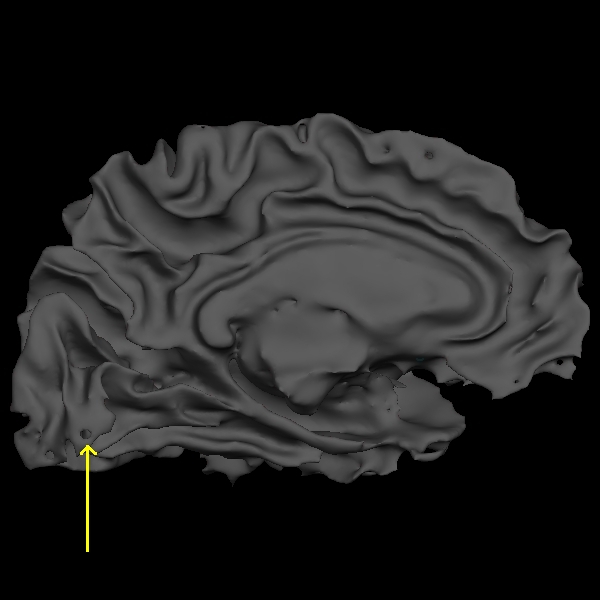
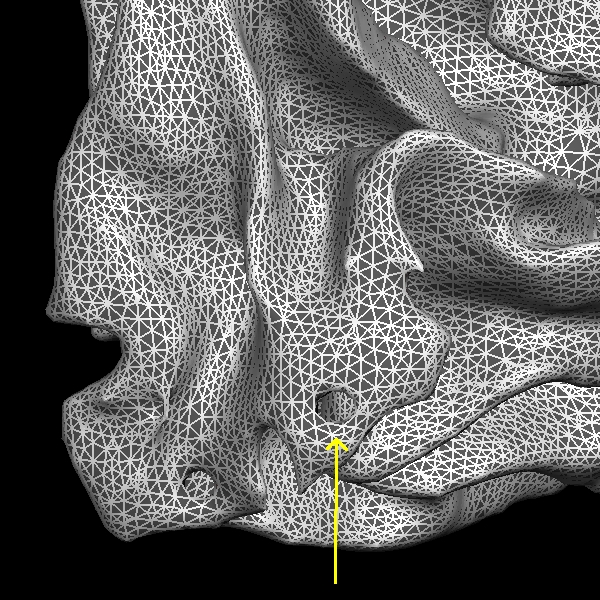
Back in the coronal view, if you scroll through slice by slice, you'll notice that there is a small group of white matter voxels which disconnect and connect back again after a few slices. The white matter surface doesn't include these voxels when they are disconnected. This area is too thin to add control points to extend the WM surface. You'll need to add some white matter voxels to fix this error. But first scroll through the slices in the brainmask.mgz volume and make a note of where the white surface is just beginning to exclude voxels and where the problem ends. In this case, the problem starts at coronal slice 53 and goes on up to slice 60.
To begin editing, click on the Recon Edit button on the freeview tool bar. In the Recon Edit window that pops up, you'll see that the the Recon editing checkbox is selected, setting the brush value to 255 and eraser value to 1. Double-check that the wm volume is on top of the brainmask and highlighted. Your viewing window should look like this:
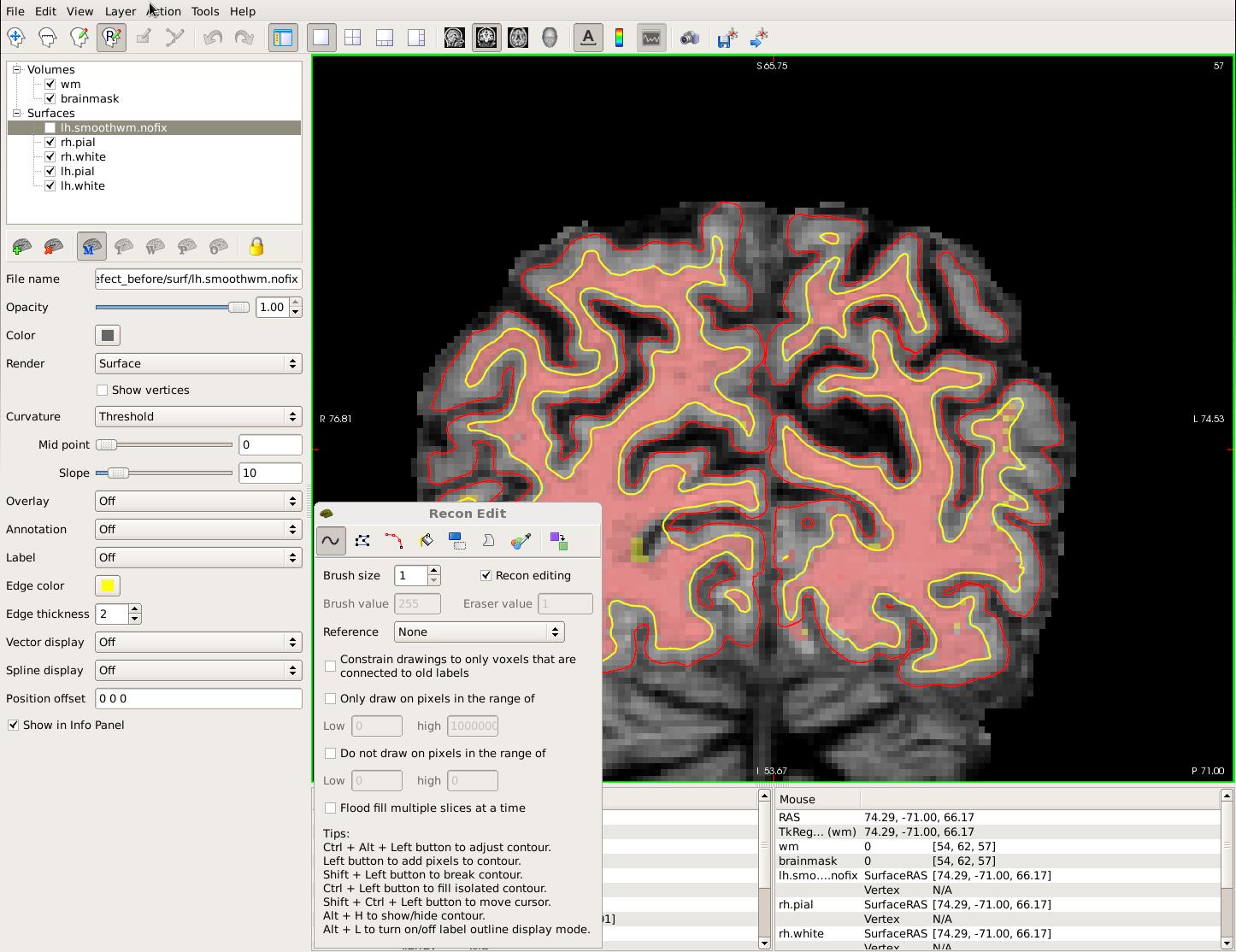
If the wm.mgz is not on top and highlighted in the Volumes section of the sidebar, you will be editing the brainmask.mgz. Use the left mouse button to begin painting in the voxels. Hold down the Shift key while clicking with the left mouse button to delete voxels.
Start adding the voxels to the wm.mgz volume on coronal slice 53. Toggle between the brainmask.mgz and wm.mgz by unchecking the check box next to the wm.mgz in the volumes tab or using Alt-V while wm.mgz is highlighted to hide and unhide it. Use brainmask.mgz as a guide while painting. Only add as many voxels as is needed to correct the error. Switch between the coronal, sagittal and horizontal views to check if you need to add more for the white matter to be labelled correctly. With more practice, it becomes easier to decide where to add voxels to correct the defect.
Video Tutorial:
At any time, you can save the changes you've made to the wm volume (for the sake of the course, do not press save) by selecting the Save Volume button  or going to File --> Save Volume while the wm.mgz is highlighted. You can look at wm.mgz volume of topo_defect_after to compare your edits with the tutorial data.
or going to File --> Save Volume while the wm.mgz is highlighted. You can look at wm.mgz volume of topo_defect_after to compare your edits with the tutorial data.
freeview -v topo_defect_after/mri/brainmask.mgz \ topo_defect_after/mri/wm.mgz:colormap=heat:opacity=0.4:visible=0 \ -f topo_defect_after/surf/lh.white:edgecolor=yellow \ topo_defect_after/surf/lh.pial:edgecolor=red \ topo_defect_after/surf/rh.white:edgecolor=yellow \ topo_defect_after/surf/rh.pial:edgecolor=red \ topo_defect_after/surf/lh.orig.nofix:visible=0
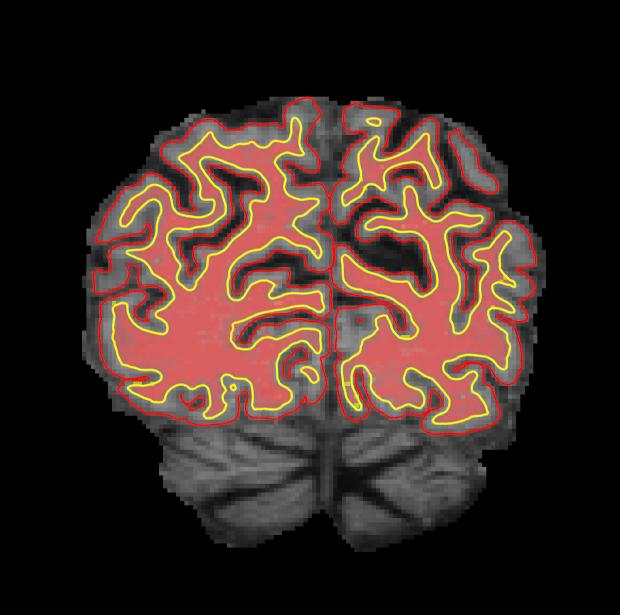
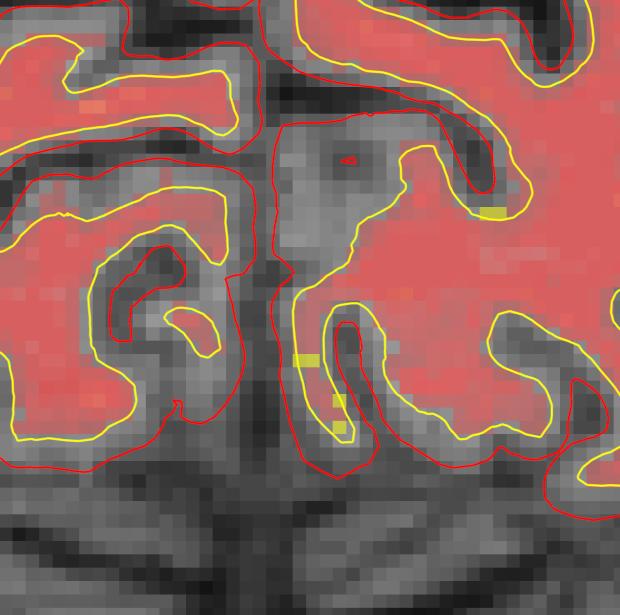
Coronal slice 57 in brainmask.mgz looks like this after correction. The yellow voxels in the zoomed in image are the voxels you have filled in to fix the topological error.
Regenerating the Surface
Once you have made all the edits to the wm.mgz volume, you can run the following command to regenerate the surfaces:
recon-all -autorecon2-wm -autorecon3 -subjid topo_defect_before
Do not run this command if you are conducting the tutorial!
