Troubleshooting your output
Just because recon-all finishes “without error” does not mean that everything is ok! It is recommended that you manually inspect the output of your recon. This set of exercises will take you through a few examples of problem outputs, asking you to identify the problems and possible methods to fix the problems. Each example will have a before and after picture, as well as an explanation of how to fix the problems seen. For a more in depth explanation of each type of problem, please see the course talks as well as the accompanying lecture slides here.
Preparations
If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window to paste (or use the keyboard shortcut Ctrl+Shift+v).Press enter to run the command. These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into this directory. You can now skip ahead to the tutorial (below the gray line).
If You're Not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
## bash export FREESURFER_HOME=/path/to/freesurfer source $FREESURFER_HOME/SetUpFreeSurfer.sh export TUTORIAL_DATA=<path_to_your_tutorial_data> export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR ## tcsh setenv FREESURFER_HOME /path/to/freesurfer source $FREESURFER_HOME/SetUpFreeSurfer.csh setenv TUTORIAL_DATA <path_to_your_tutorial_data> setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR
If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
Warming up
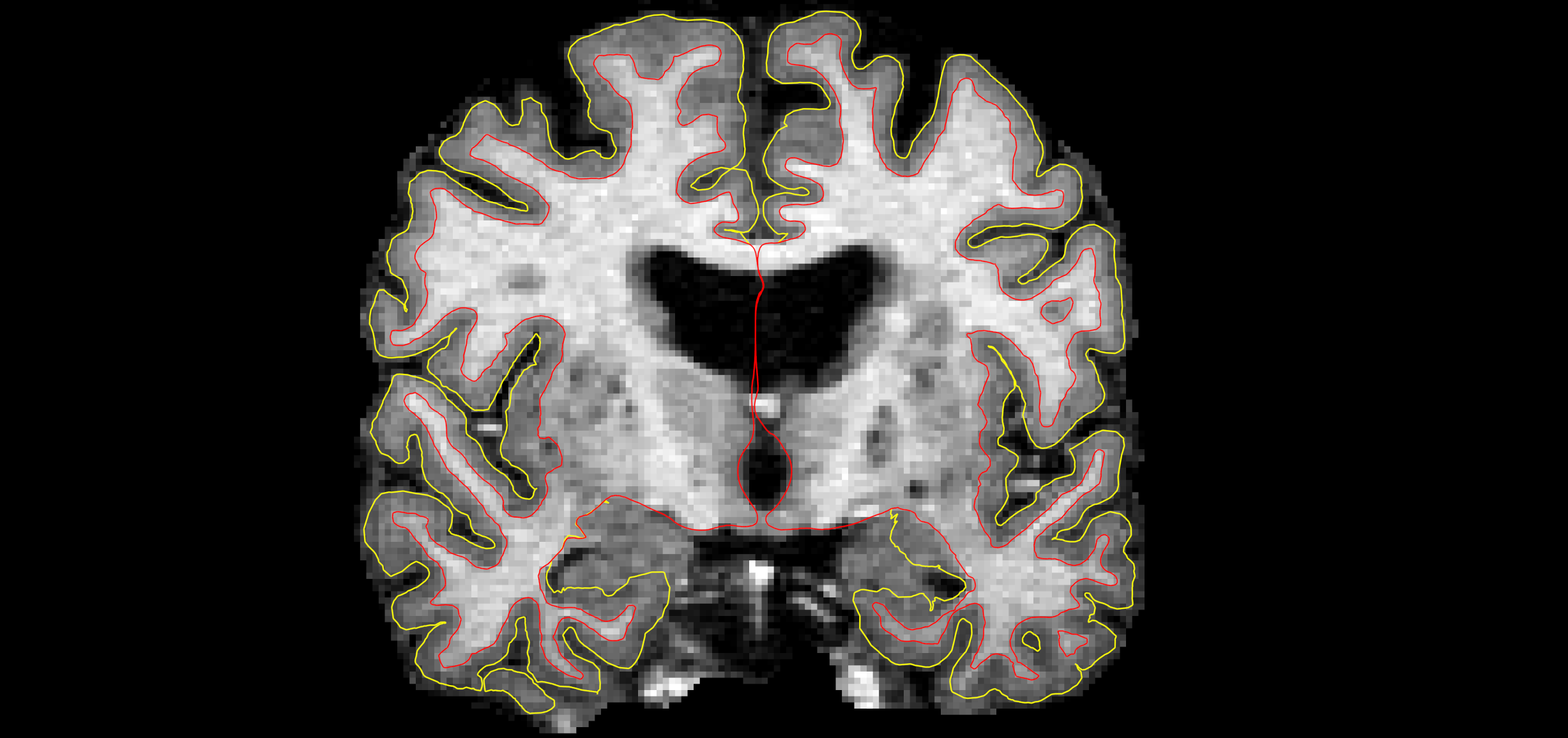
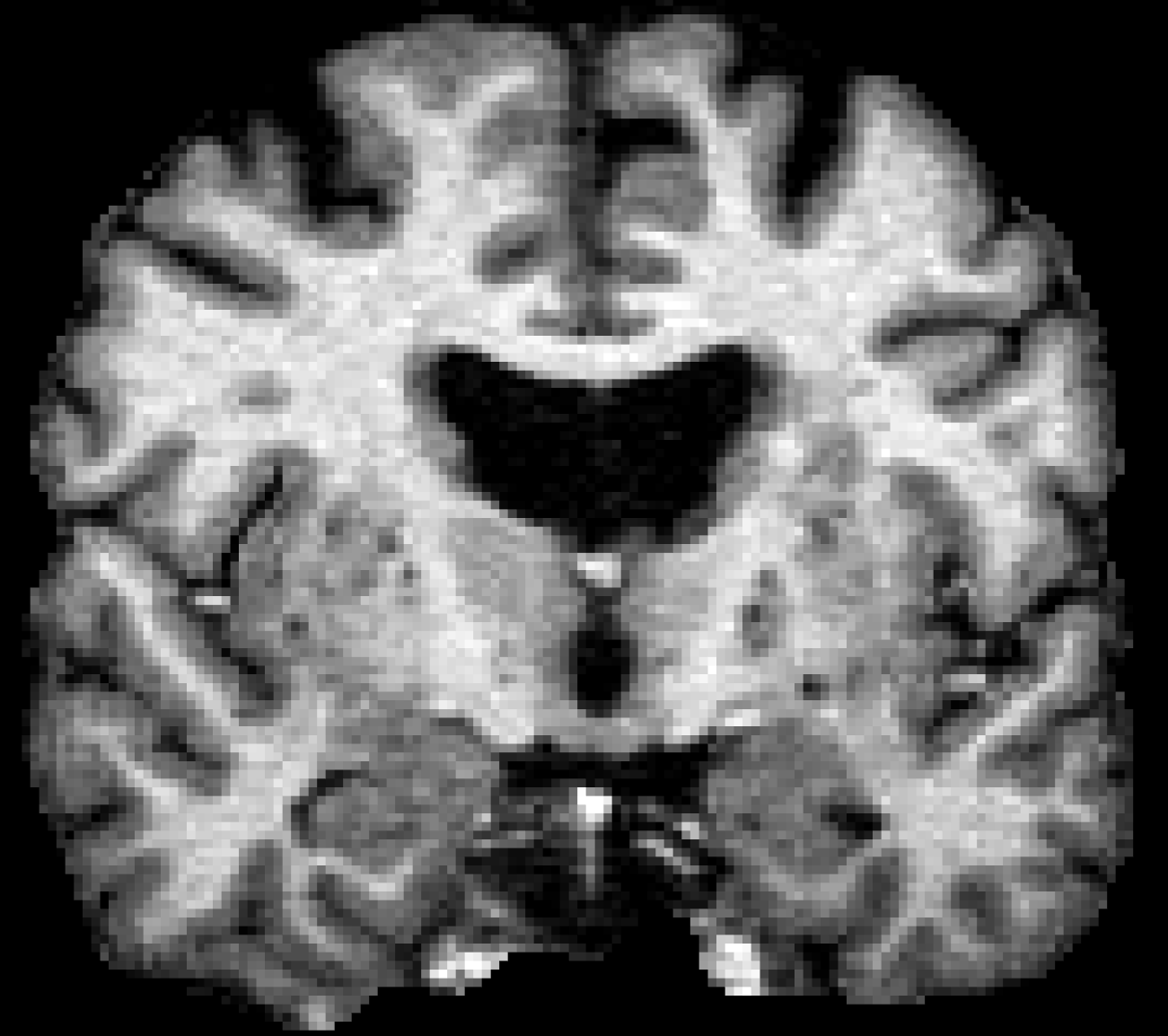
To get yourself in the right frame of mind to check for surface errors, complete the following exercise. Below is a slice of volume data. Your goal is to try and draw the surfaces yourself. First, trace with your mouse or a pen (either on paper or in an image editor) along where you think the boundary between white and grey is (the white matter surface). Then, trace along where you think the pial surface is (between grey and cerebral-spinal fluid). While you trace circle any areas you are not sure about.
Don't worry about getting things perfect! This exercise is just to help you think about and look at surfaces. When done, the surfaces as identified by FreeSurfer are shown at the very bottom of this page. Scroll down, see how those results compare to yours. Be sure to compare your circled areas. Then scroll back up to the next section to continue.
Identifying errors (a review)
Below, we ask you to look at several subjects with various errors that require manual intervention. Open each subject and try to identify the problem on your own. For each subject, there is a link to instructions on how to fix the problem. If you need a reminder of how to view the various outputs, you can refer back to the previous exercise. Errors occur when recon-all finishes but the white or pial surfaces (or less frequently, the aseg) are inaccurate. When they are inaccurate, you have to manually change certain files created during the recon-all processing stream and then regenerate the surfaces. It is not possible to directly edit the location of a surface. To begin checking the recon for accuracy, first ask yourself these two questions:
- Do the surfaces follow gray matter/white matter borders?
- Does the subcortical segmentation follow intensity boundaries?
If the answer to either is 'no', you will have to manually edit your subject. There are five main types of errors/recon-all failures, and four main ways to fix these errors. The types of errors are:
- Skull Strip Errors
- Segmentation Errors
- Intensity Normalization Error
- Pial Surface misplacement
- Topological Defect
And ways to fix these errors:
- Erase voxels
- Fill voxels
- Clone voxels (ie, copy from one volume to another)
- Add “Control Points”
To see a cheat sheet for figuring out which edits to make, please see the table under Tips and general questions.
Fixing errors
Edits to the brainmask volume (pial surface error)
Open the subject, pial_edits_before.
freeview -v pial_edits_before/mri/T1.mgz \ pial_edits_before/mri/brainmask.mgz \ -f pial_edits_before/surf/lh.white:edgecolor=yellow \ pial_edits_before/surf/lh.pial:edgecolor=red \ pial_edits_before/surf/rh.white:edgecolor=yellow \ pial_edits_before/surf/rh.pial:edgecolor=red
NOTE: The backslash (\) in the command above treats multiple lines of commands as if it is all on one line. We use this throughout the tutorials to display the commands in a more easy-to-read manner, while still allowing you to copy and paste. The backslash is not required to make the command work.
The command above will open the brainmask.mgz and T1.mgz volumes and the surfaces for both hemispheres. Scroll through the slices of this subject to check the pial surface for accuracy, making sure it follows the actual graymatter/CSF boundaries and that it does not include anything that should not be included. Once you have identified the problem (or have given up!), click here for detailed instructions on how you can fix it using Freeview. You can also compare the original data to a corrected and rerun version, pial_edits_after. To view this corrected subject (which was corrected by following the detailed instructions provided to you):
freeview -v pial_edits_after/mri/T1.mgz \ pial_edits_after/mri/brainmask.mgz \ -f pial_edits_after/surf/lh.white:edgecolor=yellow \ pial_edits_after/surf/lh.pial:edgecolor=red \ pial_edits_after/surf/rh.white:edgecolor=yellow \ pial_edits_after/surf/rh.pial:edgecolor=red
Edits to the wm volume (segmentation error)
Now, take a look at the next subject, wm1_edits_before.
freeview -v wm1_edits_before/mri/brainmask.mgz \ wm1_edits_before/mri/wm.mgz:colormap=heat:opacity=0.4 \ -f wm1_edits_before/surf/lh.white:edgecolor=yellow \ wm1_edits_before/surf/lh.pial:edgecolor=red \ wm1_edits_before/surf/rh.white:edgecolor=yellow \ wm1_edits_before/surf/rh.pial:edgecolor=red \ wm1_edits_before/surf/rh.inflated:visible=0 \ wm1_edits_before/surf/lh.inflated:visible=0
Again, this will bring up the brainmask, the wm.mgz volume, the surfaces for both hemispheres, as well as the inflated surfaces. The trouble with this subject has occurred during the white matter segmentation step. Check the white surface to find the spot that does not match the actual gray/white boundary. In the wm.mgz volume you will see a hole in this area. Click here for detailed instructions on how you can fix it using Freeview. You can also compare the original data to a corrected and rerun version, wm1_edits_after. To view this corrected subject (which was corrected by following the detailed instructions provided to you):
freeview -v wm1_edits_after/mri/T1.mgz \ wm1_edits_after/mri/brainmask.mgz \ -f wm1_edits_after/surf/lh.white:edgecolor=yellow \ wm1_edits_after/surf/lh.pial:edgecolor=red \ wm1_edits_after/surf/rh.white:edgecolor=yellow \ wm1_edits_after/surf/rh.pial:edgecolor=red
Correcting topological defects (white surface error)
Open the subject, topo_defect_before, in freeview.
freeview -v topo_defect_before/mri/brainmask.mgz \ topo_defect_before/mri/wm.mgz:colormap=heat:opacity=0.4 \ -f topo_defect_before/surf/lh.white:edgecolor=yellow \ topo_defect_before/surf/lh.pial:edgecolor=red \ topo_defect_before/surf/rh.white:edgecolor=yellow \ topo_defect_before/surf/rh.pial:edgecolor=red
This will open the brainmask.mgz volume, the wm.mgz, and the surfaces for both hemispheres. Take a look at this subject and see if you can identify what is wrong. Hint: It's a very small error in the posterior part of the brain. Once you spot the error, click here for detailed instructions on how you can fix it using Freeview. After you have fixed it, you can compare the original data to the corrected and rerun version, topo_defect_after. To view this corrected subject:
freeview -v topo_defect_after/mri/brainmask.mgz \ topo_defect_after/mri/wm.mgz:colormap=heat:opacity=0.4 \ -f topo_defect_after/surf/lh.white:edgecolor=yellow \ topo_defect_after/surf/lh.pial:edgecolor=red \ topo_defect_after/surf/rh.white:edgecolor=yellow \ topo_defect_after/surf/rh.pial:edgecolor=red
Skull strip
Now, take a look at the next subject, skullstrip_before, in freeview.
freeview -v skullstrip_before/mri/T1.mgz \ skullstrip_before/mri/brainmask.mgz:visible=false \ -f skullstrip_before/surf/lh.white:edgecolor=yellow \ skullstrip_before/surf/lh.pial:edgecolor=red \ skullstrip_before/surf/rh.white:edgecolor=yellow \ skullstrip_before/surf/rh.pial:edgecolor=red
The trouble with this subject has occurred in the skull stripping step. Check the brainmask.mgz volume carefully, comparing it to the T1.mgz volume to make sure that the skull has been completely stripped away, leaving behind the complete cortex and the cerebellum. Click here for detailed instructions on how you can fix it using Freeview. You can also compare this to the finished version, skullstrip_after. To view this corrected subject:
freeview -v skullstrip_after/mri/T1.mgz \ skullstrip_after/mri/brainmask.mgz \ -f skullstrip_after/surf/lh.white:edgecolor=yellow \ skullstrip_after/surf/lh.pial:edgecolor=red \ skullstrip_after/surf/rh.white:edgecolor=yellow \ skullstrip_after/surf/rh.pial:edgecolor=red
Adding control points (intensity normalization error)
Now, take a look at the next subject, cp_before.
freeview -v cp_before/mri/brainmask.mgz \ cp_before/mri/T1.mgz \ -f cp_before/surf/lh.white:edgecolor=blue \ cp_before/surf/lh.pial:edgecolor=red \ cp_before/surf/rh.white:edgecolor=blue \ cp_before/surf/rh.pial:edgecolor=red
The trouble with this subject has occurred during the intensity normalization. Check the white surface and be sure it is including all of the white matter as it should. If you find an area that is not included as white matter be sure to check to see that the brainmask voxel intensity is at (or very close to) 110. Click here for detailed instructions on how you can fix it using Freeview. You can also compare the original data to a corrected and rerun version, cp_after. To view this corrected subject (which was corrected by following the detailed instructions provided to you):
freeview -v cp_after/mri/brainmask.mgz \ cp_after/mri/T1.mgz \ -f cp_after/surf/lh.white:edgecolor=blue \ cp_after/surf/lh.pial:edgecolor=red \ cp_after/surf/rh.white:edgecolor=blue \ cp_after/surf/rh.pial:edgecolor=red
Less common errors (Talairach, Aseg)
Editing the Aseg
If you are at an organized course, this is NOT something you need to work on. This section is here to refer to if you run into issues at a later point. We generally advise against editing the aseg.mgz unless absolutely necessary. Try using an expert options (xopts) script first to change the intensity thresholds being used. For more information on expert options for certain recon_all commands, see the "expert options" section of the recon-all --help documentation. Editing the aseg will add more variance to your data. Instructions on how to edit the aseg using Freeview are here.
Implementing fixes
Processing stream overview
After manual intervention, you will re-run parts of recon-all to re-generate surfaces. Below is a schematic of where each of the manual edits will affect the processing stream:
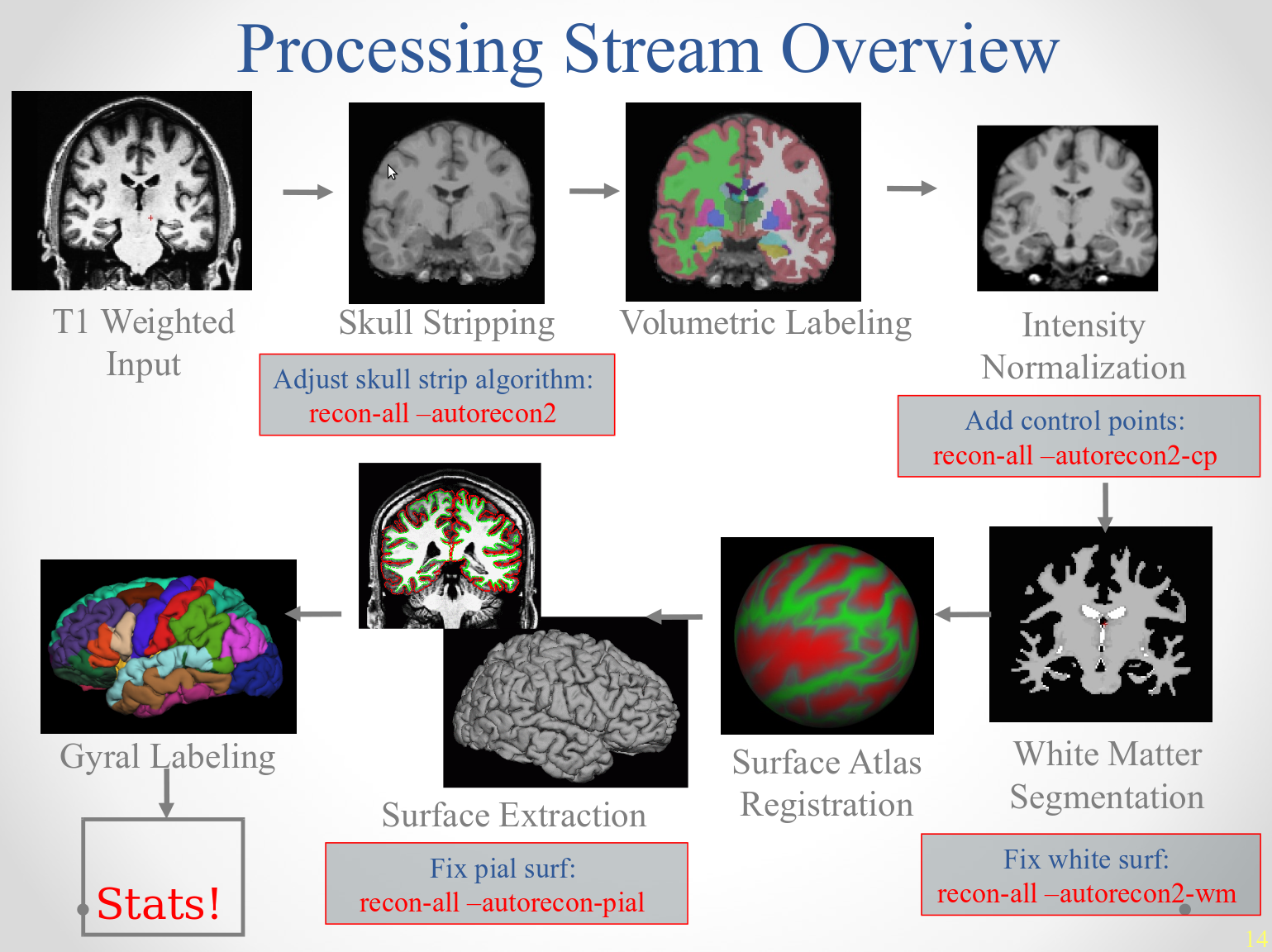
After autorecon2 flags (-autorecon2-wm, -autorecon2-cp), you must remember to also supply the -autorecon3 flag. See below for a summary of frequently used commands. Greater detail is provided the ReconAllTableStableV6.0 and ReconAllDevTable.
Frequently used commands
The following commands depend on the edits you have made to your subject output. If you've only corrected one type of error, use one of the following commands to re-run recon-all from the point in the processing stream where the error occurred. This is an efficient way of saving time when you're editing a subject.
If you've made edits to the brainmask volume and would like to re-create the pial surface, run:
recon-all -autorecon-pial -subjid <subject>
If you've made edits to the white matter volume (segmentation error or topological error) and would like to re-create final surfaces, run:
recon-all -autorecon2-wm -autorecon3 -subjid <subject>
If you've added control points:
recon-all -autorecon2-cp -autorecon3 -subjid <subject>
If you cannot remember which command to run, use the -make flag. Recon-all will automatically begin at whichever point in the processing stream you made your edits. Note: please see below under "Tips and general questions" for information on running with v6.0.
recon-all -make all -subjid <subject>
If you have corrected several types of errors, it is recommended that you re-run recon-all from the beginning, to ensure that none of your edits were left out. (Running from autorecon1 will not save you much time)
recon-all -all -subjid <subject>
Tips and general questions
Tips
- Look at all three views to determine if something is an error. Scroll back and forth a few slices – 3D structure is difficult to discern!
- All editing should take 30 minutes or less (after some practice).
- If you don't know, leave it alone.
- Don't edit too much! This reduces reliability and is almost never needed. This could cause you to need to start over (ex: too many control points, in the wrong places).
- Make a copy of your subject before editing and only make edits on this copy. That way you have the original to compare to to see if your edits worked, or you can use the original to start over again.
- If you're erasing from the brainmask volume, be careful when removing from pial surface (you don't want to accidentally erase gray matter)!
Questions
- How do you know which volumes to edit?
Answer: If the pial surface includes too much, edit brainmask.mgz. If the error affects the white surface (too much/little) or if the pial surface includes too little, edit the wm.mgz (if segmentation error) or add control points (if normalization error). This is because errors in pial surface placement are typically caused by underlying errors in the white surface placement, and can be corrected by interventions that fix the white surface. Below is a table summarizing which volumes you should be focused on editing based on your output error:
|
Non-gm in pial surf |
Non-wm in white surf |
Wm excluded from surf & intensity = 110 |
Wm excluded from surf & intensity < 110 |
Large area of brain excluded |
brainmask.mgz |
Erase |
|
|
|
Clone (from T1.mgz |
brain.finalsurfs.mgz |
Erase |
|
|
|
|
wm.mgz |
|
Erase |
Fill |
|
|
add control points (use brainmask as guide- not tied to particular volume) |
|
|
|
Control points |
|
- I made edits to my subject output and now I don't want to use them. What should I do?
Answer: To re-run recon-all without edits, use –clean flags or start from scratch.
- I fixed a topological error and the white matter surface is still incorrect. What now?
Answer: If the ?h.orig surface misses white matter that is accurately labeled in the wm.mgz or extends into regions where there is no wm in the wm.mgz, then there is an incorrectly fixed topological defect. Even one or two missing voxels can cause large-scale defects, so very minor editing (e.g. filling in white matter voxels that are holes, or erasing handles) may fix the problem.
- I ran recon-all after editing and nothing happened. What now?
Answer: Re-run recon-all using the flag -make all instead *only if you are running the FreeSurfer development version or version 5.3*. Before doing so, check to make sure your edits were saved.
recon-all -make all -s <subjectID>
If you would like to run -make all and you are using version 6.0: copy "recon-all.makefile" from the https://surfer.nmr.mgh.harvard.edu/pub/dist/freesurfer/dev_binaries site and use it to overwrite $FREESURFER_HOME/bin/recon-all.Makefile. Once this is complete you can use the make flag.
For more questions and answers, please visit the FAQ page: here
Quiz
You can test your knowledge of this tutorial by clicking here for a quiz!
First exercise surfaces