|
Size: 8761
Comment:
|
Size: 8012
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| [[Tutorials|Back to Tutorial Homepage]]<<BR>>[[FsTutorial|Back to Course Homepage]]<<BR>>[[FsTutorial/MultiModal_freeview|Back to Multimodal Tutorial List]] | [[Tutorials|Back to Tutorial Homepage]] | [[FsTutorial|Back to Course Homepage]] | [[MultiModalTutorialV6.0|Back to Multimodal Tutorial List]] |
| Line 4: | Line 4: |
| ##Other Multimodal Tutorials: [[FsTutorial/MultiModalDtiIndividual_freeview|B. Individual DTI Integration]], [[FsTutorial/MultiModalFmriIndividual_freeview|C. Individual fMRI Integration]], [[FsTutorial/MultiModalFmriGroup_freeview|D. Surface-based Group fMRI Analysis]] | |
| Line 6: | Line 5: |
| Other Multimodal Tutorials: [[FsTutorial/MultiModalFmriIndividual_freeview|Individual fMRI Integration]], [[FsTutorial/MultiModalFmriGroup_freeview|Surface-based Group fMRI Analysis]], [[FsTutorial/Diffusion_DtiIntegration|Diffusion and DTI Integration]] | Other Multimodal Tutorials: [[MultiModalTutorialV6.0/IndividualFMRI|Individual fMRI Integration]], [[MultiModalTutorialV6.0/FMRIGroupAnalysis|Surface-based Group fMRI Analysis]], [[FsTutorial/DiffusionV6.0|Diffusion and DTI Integration]] |
| Line 50: | Line 49: |
| 1. Familiarize you with the FreeView interface. | 1. Familiarize you with the !FreeView interface. |
| Line 54: | Line 53: |
Using the following command, you can run !FreeView using a wrapper script. This script can be an efficient way to open !FreeView without having to type a long, cumbersome command. |
You can run !FreeView using the following command: |
| Line 58: | Line 56: |
| tkregisterfv --mov template.nii --s fbirn-anat-101.v4 --reg manreg.lta | freeview -v template.nii \ $SUBJECTS_DIR/fbirn-anat-101.v4/mri/orig.mgz:visible=0 -f \ $SUBJECTS_DIR/fbirn-anat-101.v4/surf/lh.white \ $SUBJECTS_DIR/fbirn-anat-101.v4/surf/rh.white \ -viewport cor -transform-volume |
| Line 60: | Line 62: |
tkregisterfv will output the !FreeView command that it calls. For this example, it should output the following in the terminal: {{{ freeview -transform-volume -viewport cor -v /home/nmrclass/tutorial_data/buckner_data/tutorial_subjs/fbirn-anat-101.v4/mri/orig.mgz:visible=0:name=orig.mgz(targ) /home/nmrclass/tutorial_data/buckner_data/tutorial_subjs/multimodal/fmri/fbirn-101/template.nii:name=template.nii(mov):reg=/home/nmrclass/tutorial_data/buckner_data/tutorial_subjs/multimodal/fmri/fbirn-101/manreg.lta --surface /home/nmrclass/tutorial_data/buckner_data/tutorial_subjs/fbirn-anat-101.v4/surf/lh.white:edgecolor=yellow --surface /home/nmrclass/tutorial_data/buckner_data/tutorial_subjs/fbirn-anat-101.v4/surf/rh.white:edgecolor=yellow }}} |
|
| Line 69: | Line 64: |
| {{attachment:FreeView.Snap1.sm.png||height="448",width="717"}} | {{attachment:FreeView.Snap1.sm.png||width="717",height="448"}} |
| Line 75: | Line 70: |
| Notice how the surfaces do not line up with the anatomy in the functional volume. To see how much the registration is off, you can switch between the orig.nii volume and the template.nii volume by clicking 'template.nii(mov)' in the upper left corner and pressing Alt-c. To do manual registration in !FreeView, select the volume you want to adjust (in this case, choose template), and go to the 'Tools' menu and select 'Transform Volume' (this window should already be loaded). Select the 'Translate' tab and play around with the sliders until the surfaces are better aligned with the anatomy. You could play with the 'Rotation' sliders as well. | Notice how the surfaces do not line up with the anatomy in the functional volume. To see how much the registration is off, you can switch between the orig.nii volume and the template.nii volume by clicking 'template.nii' in the upper left corner and pressing Alt-c. To do manual registration in !FreeView, select the volume you want to adjust (in this case, choose template), and go to the 'Tools' menu and select 'Transform Volume' (this window should already be loaded). Select the 'Translate' tab and play around with the sliders until the surfaces are better aligned with the anatomy. You could play with the 'Rotation' sliders as well. |
| Line 77: | Line 72: |
| {{attachment:FreeView.Snap2.png||height="410",width="386"}} {{attachment:FreeView.Snap3.png||height="410",width="386"}} | {{attachment:FreeView.Snap2.png||width="386",height="410"}} {{attachment:FreeView.Snap3.png||width="386",height="410"}} |
| Line 79: | Line 74: |
| When you are happy with your results, save the registration file ('Save Reg') and/or the resampled volume ('Save Volume As'). If you would like to start over, you can choose 'Restore to Original'. Note: !FreeView will save .lta format registration files. | When you are happy with your results, save the registration file ('Save Reg') and/or the resampled volume ('Save Volume As') as either register_manual or register_<date>. If you would like to start over, you can choose 'Restore to Original'. Note: !FreeView will save .lta format registration files. |
| Line 85: | Line 80: |
| bbregister --mov template.nii --bold --s fbirn-anat-101.v4 --reg register.lta | bbregister --mov template.nii --bold --s fbirn-anat-101.v4 --lta register.lta |
| Line 99: | Line 94: |
| Click [[FsTutorial/MultiModalRegistration/RegisterFileLta|here]] to see an example. | Click [[MultiModalTutorialV6.0/MultiModalRegistration/RegisterFileLta|here]] to see an example. |
| Line 106: | Line 101: |
| Line 112: | Line 106: |
| tkregisterfv --mov template.nii --reg register.lta --surfs | freeview -v $SUBJECTS_DIR/fbirn-anat-101.v4/mri/orig.mgz:visible=0 \ template.nii:reg=register.lta -f \ $SUBJECTS_DIR/fbirn-anat-101.v4/surf/lh.white \ $SUBJECTS_DIR/fbirn-anat-101.v4/surf/rh.white \ -viewport cor -transform-volume |
| Line 114: | Line 112: |
| Select the template volume and hit Alt+c to switch between the template and orig, or change the opacity value on the top volume to observe the quality of the alignment. See how much better the registration is now? How does it compare to your manual registration? Note that you do not need to specify the subject name because that is in the registration file generated by bbregister. | Select the template volume and hit Alt+c to switch between the template and orig, or change the opacity value on the top volume to observe the quality of the alignment. How does this registration look compared to your manual registration above? Can you make it any better? |
| Line 116: | Line 114: |
| {{attachment:Registered.Snap.sm.png||height="428",width="684"}} | {{attachment:Registered.Snap.sm.png||width="684",height="428"}} |
| Line 118: | Line 116: |
| Notes: | ---- == Study Question == *Which imaging plane(s) (axial/horizontal, sagittal, coronal) move on on the x axis when you drag the X (L-R) Translate bar? [[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/StudyQuestions#Whichimagingplane.28s.29.28axial.2BAC8-horizontal.2Csagittal.2Ccoronal.29moveononthexaxiswhenyoudragtheX.28L-R.29Translatebar.3F|Answer]] |
| Line 120: | Line 120: |
| 1. The subject is not specified since it is in the register.dat. 1. --regheader is not used. |
----- |
| Line 123: | Line 122: |
| How does this registration look compared to your manual registration above? Can you make it any better? | |
| Line 127: | Line 125: |
| Other Multimodal Tutorials: [[FsTutorial/MultiModalFmriIndividual_freeview|Individual fMRI Integration]], [[FsTutorial/MultiModalFmriGroup_freeview|Surface-based Group fMRI Analysis]], [[FsTutorial/Diffusion_DtiIntegration|Diffusion and DTI Integration]] | Other Multimodal Tutorials: [[MultiModalTutorialV6.0/IndividualFMRI|Individual fMRI Integration]], [[MultiModalTutorialV6.0/FMRIGroupAnalysis|Surface-based Group fMRI Analysis]], [[FsTutorial/DiffusionV6.0|Diffusion and DTI Integration]] |
| Line 129: | Line 127: |
| <<BR>>[[Tutorials|Back to Tutorial Homepage]]<<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial|Back to Course Homepage]] | <<BR>>[[Tutorials|Back to Tutorial Homepage]] <<BR>>[[https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial|Back to Course Homepage]] |
Back to Tutorial Homepage | Back to Course Homepage | Back to Multimodal Tutorial List
Other Multimodal Tutorials: Individual fMRI Integration, Surface-based Group fMRI Analysis, Diffusion and DTI Integration
Registration
The purpose of this multimodal tutorial is to get you acquainted with the concepts needed to perform multi-modal integration in FreeSurfer using fMRI and DTI analysis. You will not learn how to perform fMRI or DTI analysis here; that knowledge is already assumed. The fMRI makes use of data from the Functional Biomedical Informatics Research Network (fBIRN, www.nbirn.net).
Contents
1. Preparations
1.1. If You're at an Organized Course
If you are taking one of the formally organized courses, everything has been set up for you on the provided laptop. The only thing you will need to do is run the following commands in every new terminal window (aka shell) you open throughout this tutorial. Copy and paste the commands below to get started:
export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101
To copy: Highlight the command in the box above, right click and select copy (or use keyboard shortcut Ctrl+c), then use the middle button of your mouse to click inside the terminal window (this will paste the command). Press enter to run the command.
These two commands set the SUBJECTS_DIR variable to the directory where the data is stored and then navigates into the directory within the SUBJECTS_DIR that you will use for this part of the tutorial. You can now skip ahead to the tutorial (below the gray line).
1.2. If You're Not at an Organized Course
If you are NOT taking one of the formally organized courses, then to follow this exercise exactly be sure you've downloaded the tutorial data set before you begin. If you choose not to download the data set you can follow these instructions on your own data, but you will have to substitute your own specific paths and subject names. These are the commands that you need to run before getting started:
## bash <source_freesurfer> export TUTORIAL_DATA=<path_to_your_tutorial_data> export SUBJECTS_DIR=$TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101 ## tcsh source $FREESURFER_HOME/SetUpFreeSurfer.csh setenv TUTORIAL_DATA <path_to_your_tutorial_data> setenv SUBJECTS_DIR $TUTORIAL_DATA/buckner_data/tutorial_subjs cd $SUBJECTS_DIR/multimodal/fmri/fbirn-101
Information on how to source FreeSurfer is located here.
If you are not using the tutorial data you should set your SUBJECTS_DIR to the directory in which the recon(s) of the subject(s) you will use for this tutorial are located.
2. Basic Manual Registration
The purpose of this exercise is to:
Familiarize you with the FreeView interface.
- Show you how to make manual adjustments to the registration.
- Show you what a bad registration is and how hard it is to fix manually.
You can run FreeView using the following command:
freeview -v template.nii \
$SUBJECTS_DIR/fbirn-anat-101.v4/mri/orig.mgz:visible=0 -f \
$SUBJECTS_DIR/fbirn-anat-101.v4/surf/lh.white \
$SUBJECTS_DIR/fbirn-anat-101.v4/surf/rh.white \
-viewport cor -transform-volumeIt should also bring up an interface that looks like this:
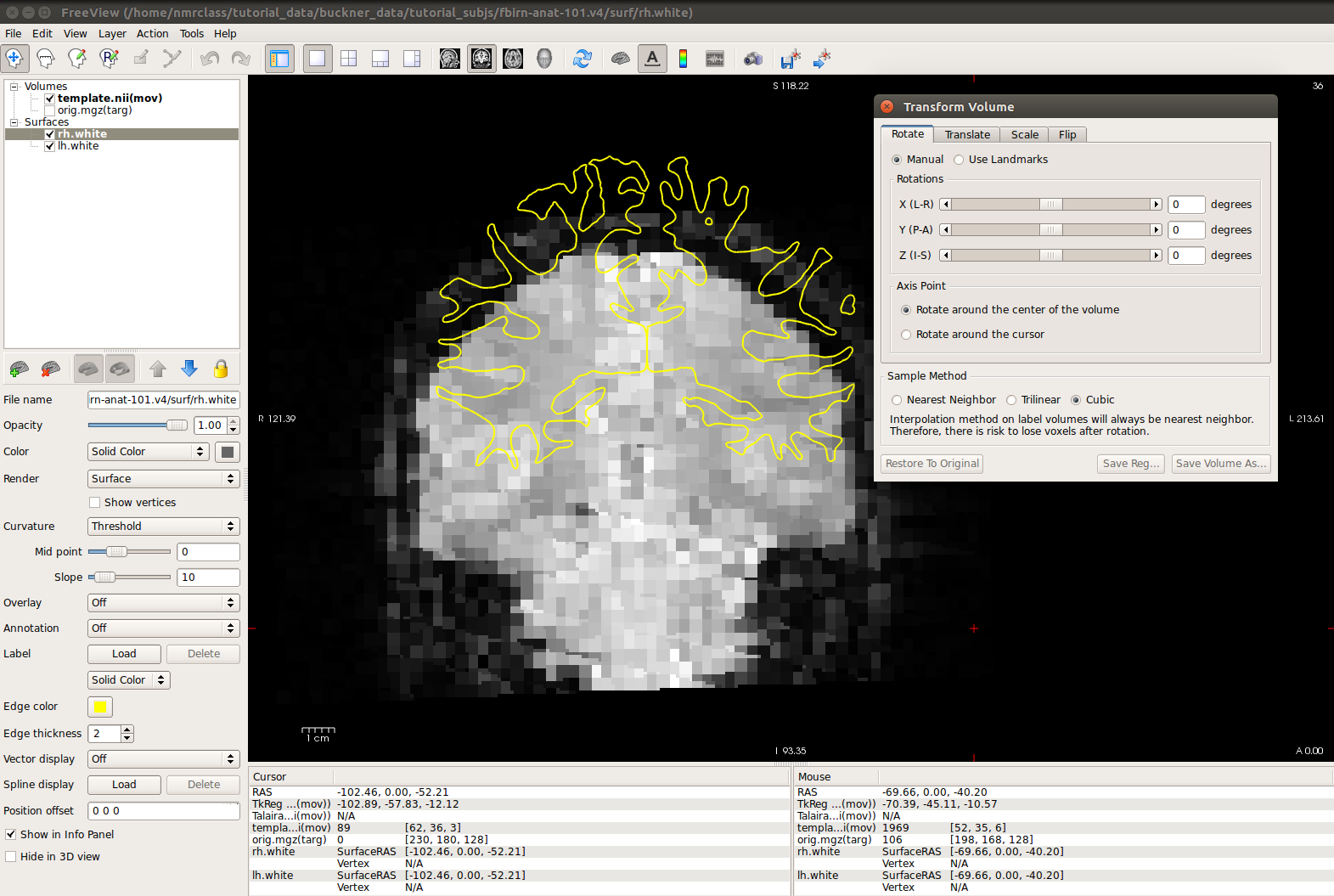
- The grayscale image is the functional data.
- The yellow line is the white surface.
- Gray matter and CSF are bright while white matter is dark.
Notice how the surfaces do not line up with the anatomy in the functional volume. To see how much the registration is off, you can switch between the orig.nii volume and the template.nii volume by clicking 'template.nii' in the upper left corner and pressing Alt-c. To do manual registration in FreeView, select the volume you want to adjust (in this case, choose template), and go to the 'Tools' menu and select 'Transform Volume' (this window should already be loaded). Select the 'Translate' tab and play around with the sliders until the surfaces are better aligned with the anatomy. You could play with the 'Rotation' sliders as well.
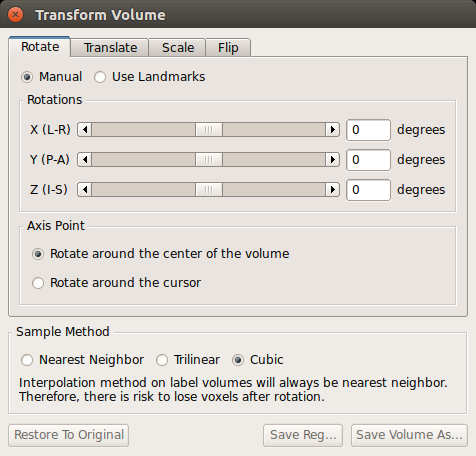
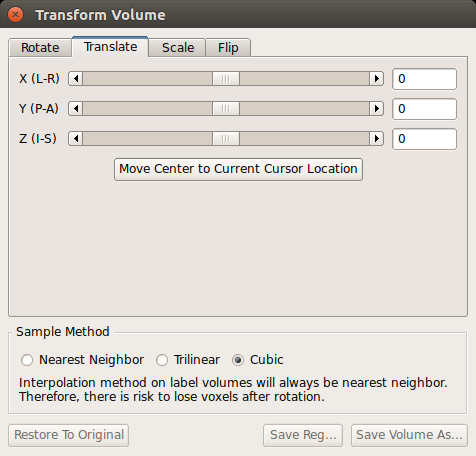
When you are happy with your results, save the registration file ('Save Reg') and/or the resampled volume ('Save Volume As') as either register_manual or register_<date>. If you would like to start over, you can choose 'Restore to Original'. Note: FreeView will save .lta format registration files.
3. Automatic Registration
The purpose of this exercise is to show you how to use the automatic registration program (bbregister). To run, copy and paste this command into your shell:
bbregister --mov template.nii --bold --s fbirn-anat-101.v4 --lta register.lta
Notes:
- --bold indicates that the template has bold/T2 contrast (meaning that gray matter is brighter than white matter).
- --lta indicates that the output registration file is register.lta (Note: currently, register.dat will also be written into the output directory for backwards compatibility).
- This will take about 5 min (the same amount of time you spent manually, right?).
4. View Automatic Registration
Look at the register.lta text file created:
cat register.lta
Click here to see an example.
You can also look at the register.dat.mincost file:
cat register.dat.mincost
The mincost file consists of 4 values. The first value indicates the quality of the registration on a scale from 0 (perfectly registered) to 1 (not registered).
In order to load a functional volume with the anatomical volume specifying the recently created automatic registration:
freeview -v $SUBJECTS_DIR/fbirn-anat-101.v4/mri/orig.mgz:visible=0 \
template.nii:reg=register.lta -f \
$SUBJECTS_DIR/fbirn-anat-101.v4/surf/lh.white \
$SUBJECTS_DIR/fbirn-anat-101.v4/surf/rh.white \
-viewport cor -transform-volumeSelect the template volume and hit Alt+c to switch between the template and orig, or change the opacity value on the top volume to observe the quality of the alignment. How does this registration look compared to your manual registration above? Can you make it any better?
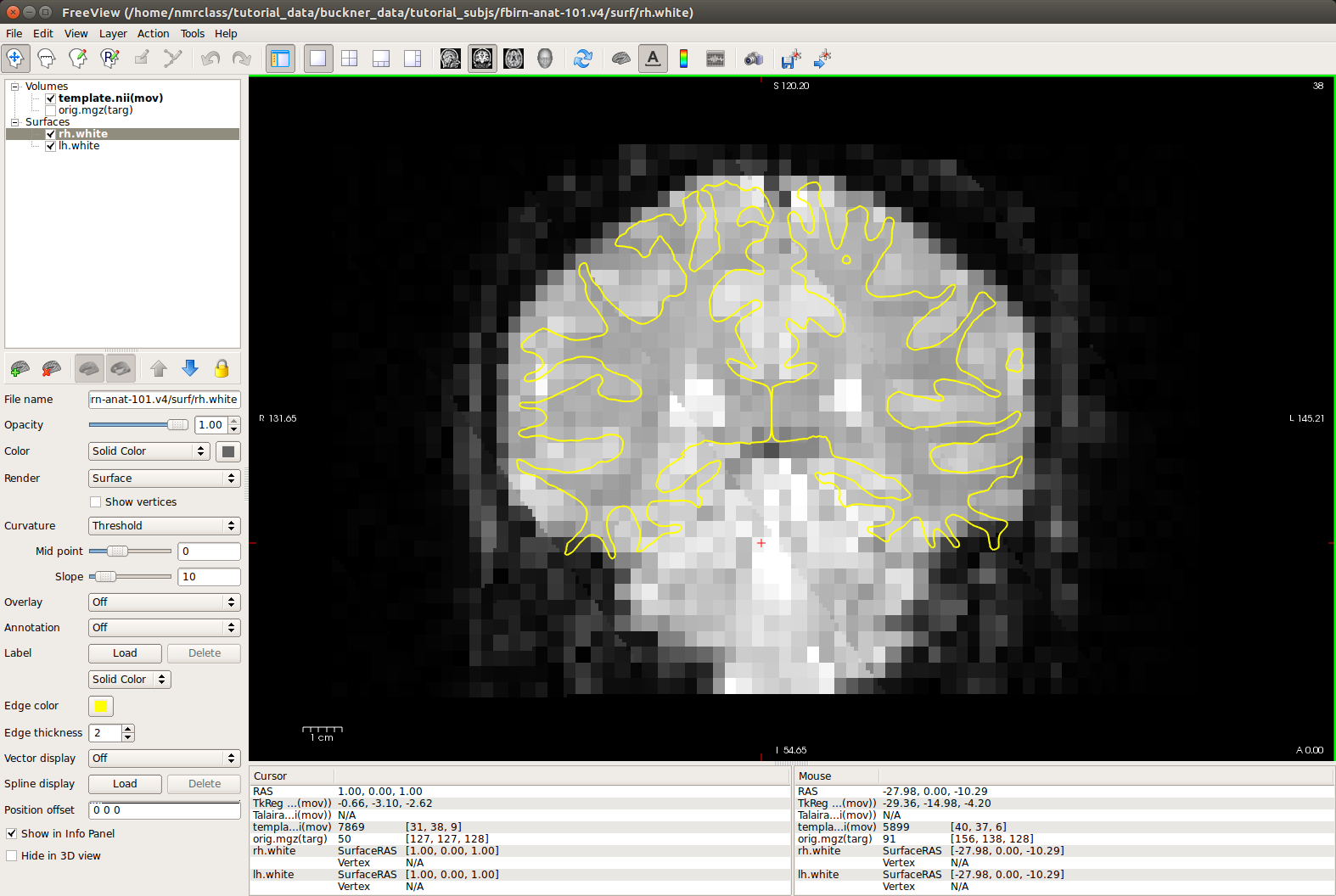
5. Study Question
Which imaging plane(s) (axial/horizontal, sagittal, coronal) move on on the x axis when you drag the X (L-R) Translate bar? Answer
Other Multimodal Tutorials: Individual fMRI Integration, Surface-based Group fMRI Analysis, Diffusion and DTI Integration
